Abstract
Contamination with fungi and their toxins is considered one of the most dangerous hidden pollutants that threaten the health of the consumer. The presence of mycotoxins in various foods has been recorded, despite their apparent safety for human consumption. The current study was conducted to evaluate the prevalence of Aspergillus species and aflatoxin producing genes using culture and molecular methods, respectively. A total of 75 samples of chicken cuts, represented by wing, breast, and thigh (25 of each), were randomly collected from various grocery and poultry shops located at Gharbiya governorate, Egypt. Results of culture and isolation techniques revealed the detection of Aspergillus sp. in 36, 48 and 44% of the examined wing, breast and thigh samples, respectively. Moreover, microbiological identification of the isolated strains showed presence of A. niger, A. flavus, A. fumigatus, A. terreus and A. parasiticus in 16, 13.3, 10.6, 1.3 and 1.3% of the total population of the examined samples. Molecular detection of some aflatoxin production regulating genes (OmtA, Nor1 and Ver1) in ten Aspergillus sp. isolates revealed their detection in 8/10 (80%), 8/10 (80%) and 7/10 (70%) represented by positive bands at molecular weight of 1024, 400 and 537 bp, respectively. Referring to the recorded results, chicken cuts may possess a great silent hazard to human being under improper good manufacturing practices and inadequate hygienic conditions during handling and storage.
Keywords
Aspergillus species; Chicken meat cuts; cPCR; Egypt
Introduction
The production of chicken meat and its derived products in developing countries plays an essential role in supporting food security and satisfying poultry meat demand [1].
The contamination of meat products with molds can occur during different preparation stages during slaughtering under bad hygienic conditions using contaminated water or by adding contaminated spices with mold spores or during packing, handling, transportation and storage [2].
The contamination of meat with Aspergillus section flavi is one of the most hazardous microbial contaminations as most species of this section are able to produce aflatoxins that can cause severe diseases and has carcinogenic effects [3].
Acute aflatoxin poisoning may lead to death as was recorded in Kenya in 2004 [4], whereas chronic poisoning may lead to various recorded mutagens and cancers [5].
Depending on their toxigenic impacts on food and human health, Aspergillus sp. is classified into two groups: the aflatoxigenic species such as A. flavusand A. parasiticus, and the non-aflatoxigenic strains such as A. tamarii and A. oryzae [6].
Molecular analyses have been used to confirm aflatoxin productivity of Aspergillus species isolates. omtA, nor1 and ver1 genes are from the commonly used genes encoded aflP, aflD and aflM toxins detection in food items [7] yield an accurate, rapid and reliable records of toxigenic Aspergillus species especially in food chain [8].
Therefore, the main target of the current study was to investigate the presence of toxigenic Aspergillus species in some chicken meat cuts collected from Gharbiya Governorate markets, Egypt.
Materials and Methods
Collection of samples
A total of seventy-five random samples of raw chilled chicken wing, thigh, and breast (25 of each) were collected from different local poultry shops and different supermarkets at Gharbia Governorate, Egypt. Samples were taken aseptically in polyethylene bags and were transferred to the laboratory in an ice box for mycological examination.
Preparation of samples [9]
Twenty-five grams from each sample were carefully and aseptically homogenized in blinder after mixing with 225 ml of sterile peptone water 0.1% to form a dilution of 1:10, from which 10-fold serial dilutions were prepared.
Determination of Aspergillus species
Culture of the prepared samples: Culture of the prepared samples was performed according to ISO [10], where 0.1ml of the previously prepared serial dilutions was spread by means of sterile L-shape glass rod over two Petri-dishes containing solidified Dichloran Rose Bengal Agar with Chloramphenicol (DRBC) were then incubated at upright position at 25°C for 5 -7 days.
Identification of isolated strains: Identification of isolated strains was performed according to Pitt JI, et al. [11] macroscopically and microscopically as recorded in table 1.
|
Colony Diameter (mm) |
Texure |
Surface color |
Reverse color |
Stipes |
Vesicles |
Serriation |
Conidia |
Colonial head/ cleistotheca |
|
A. flavus |
65-70 |
Floccose powdery or granular |
Greenish yellow |
Pale brown |
Rough hyaline |
Globose or subglobose |
Biseriate |
Globose to ellipsoid |
Radiating head |
|
A. fumaiguts |
40-70 |
Vevety to powder |
Blue with white margin |
Slight green |
Smooth hyaline |
Clavate |
Uniseriate |
Globose or subglobose |
Columner head |
| A. nidulans |
50-65 |
Velvety |
Green |
Brown |
Smooth brown |
Pyriform |
Biseriate |
Globose rough |
Radiate head/ hulla Cells ascs |
| A. niger |
50-70 |
Granular of powdery |
Black |
Pale yellow |
Smooth Yellow to brown |
Round |
Biseriate |
Globose brown |
Round head |
| A. terus |
40-60 |
Powdery |
Sandy to brown |
Pale brown |
Smooth hyaline |
Round to pyriform |
Biseriate |
Globose to ellipsiod |
Columner head |
Table 1: Morphological character of aspergillus species [11].
Molecular detection of some aflatoxin producing genes of some isolated Aspergillus strains by cPCR
Oligonucleotide primers used in cPCR: Three pairs of omtA, nor1 and ver1 primers were prepared and collected from Metabion (Germany). Their special sequence and amplify certain products as were be displayed in table 2.
| Gene |
Sequence (5'----3') |
Amplified product (bp) |
Reference |
|
omtA |
GGCCCGGTTCCTTGGCTCCTAAGC |
1024 |
Norlia M, et al. 2019 [29] |
|
|
nor1 |
ACCGCTACGCCGGCACTCTCGGCAC |
400 |
|
|
ver1 |
GCCGCAGGCCGCGGAGAAAGTGGT |
537 |
|
Table 2: Oligonucleotide primers sequences.
Mycological DNA was extracted following QIAamp DNeasy Plant Mini kit Catalogue no. 69104.
Preparation of master mix and thermal profile was adapted according to the manufacturer instructions (Emerald Amp GT PCR master mix (Takara) Code No. RR310A).
Results
As recorded in table 3, Aspergillus sp. was detected in 32 (42.6%) of the total examined samples. In detail, breast samples recorded the highest contamination level (48%); followed by thigh and wing samples, respectively.
|
Samples |
Wing |
Breast |
Thigh |
Total |
| No. |
%* |
No. |
%* |
No. |
%* |
No. |
%** |
| Aspergillus sp. |
9 |
36 |
12 |
48 |
11 |
44 |
32 |
42.6 |
Table 3: Prevalence of Aspergillus species in the examined chicken meat cut samples (n= 25 of each).
%* prevalence in relation to the number of each sample (25).
%** prevalence in relation to the total number of samples population (75).
Regarding the genus identification, A. niger had the highest detection levels (16%) in the examined samples (Table 4).
| Samples |
wings |
breast |
thigh |
Total |
| Aspergillus spp. |
No. |
% |
No. |
% |
No. |
% |
No. |
% |
| A. niger |
4 |
16 |
6 |
24 |
2 |
8 |
12 |
16 |
| A. flavus |
2 |
8 |
4 |
16 |
4 |
16 |
10 |
13.3 |
| A. fumigatus |
2 |
8 |
1 |
4 |
5 |
20 |
8 |
10.6 |
| A. terreus |
1 |
4 |
- |
|
- |
|
1 |
1.3 |
| A. parasiticus |
- |
- |
1 |
4 |
- |
|
1 |
1.3 |
Table 4: Prevalence of identified Aspergillus spp. in the examined chicken meat cuts (n= 25 of each).
Referring to the obtained results of molecular detection of some aflatoxin producing genes as recorded in table 5 and figures 1-3; omtA, nor1 and ver1 genes were detected in 8/10 (80%), 8/10 (80%) and 7/10 (70%) of the examined A. flavus isolates, respectively. The presence of these genes indicated the producibility of the examined strain for aflatoxins P, D and M, respectively.

Figure 1: Agarose gel electrophoresis of cPCR of omtA (1024 bp) gene of A. flavus.
Lane L: 100 bp ladder as molecular size DNA marker.
Lane P: Control positive A. flavus for omtA gene.
Lane N: Control negative.
Lanes 1, 3, 4, 5, 6, 7, 9 and 10: Positive A. flavus for omtA gene.
Lanes 2 and 8: Negative A. flavus for omtA gene.
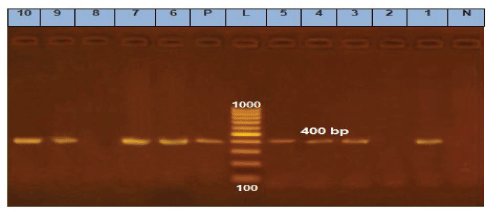
Figure 2: Agarose gel electrophoresis of cPCR of nor1 (400 bp) gene of A. flavus.
Lane L: 100 bp ladder as molecular size DNA marker.
Lane P: Control positive A. flavus for nor1 gene.
Lane N: Control negative. Lanes 1, 3, 4, 5, 6, 7, 9 and 10: Positive A. flavus for nor1gene.
Lanes 2 and 8: Negative A. flavus for nor1 gene.

Figure 3: Agarose gel electrophoresis of cPCR of ver1 (537 bp) gene of A. flavus.
Lane L: 100 bp ladder as molecular size DNA marker.
Lane P: Control positive A. flavus for ver1 gene.
Lane N: Control negative. Lanes 1, 3, 4, 5, 6, 9 and 10: Positive A. flavus for ver1 gene.
Lanes 2, 7 and 8: Negative A. flavus for ver1 gene.
| Sample |
omtA |
nor1 |
ver1 |
| 1 |
+ |
+ |
+ |
| 2 |
- |
- |
- |
| 3 |
+ |
+ |
+ |
| 4 |
+ |
+ |
+ |
| 5 |
+ |
+ |
+ |
| 6 |
+ |
+ |
+ |
| 7 |
+ |
+ |
- |
| 8 |
- |
- |
- |
| 9 |
+ |
+ |
+ |
| 10 |
+ |
+ |
+ |
Table 5: Prevalence of aflatoxin producing genes in A. flavus isolates from
the examined samples (n= 10).
Discussion
Chicken meat and meat products comply an important source of human protein supplement all over the world because they provide a good source of digestible protein, low cholesterol fat, essential amino acids, minerals, and different types of vitamins.
In Egypt, as human population increasing, demand of animal proteins also is increasing represents a serious challenge in while poultry industry plays an essential role in filling nutrition gap as a rapid and more economic source of proteins [12].
Mold contamination of meat and meat products has been considered a serious source of food spoilage, resulting in different organoleptic changes in flavor, color, texture, odor referred mainly to fungal deterioration, especially in poor developing countries due to lack of hygienic measures during processing and handling [13].
The presence of mold in foods may be referred to the rapid, easy disperse and wide spread of fungal spores, which are abundant in the environment introducing food chain through dust, water, workers and equipment. Their presence in food samples is a serious public health concern as these fungi may be associated with the production of mycotoxins [14].
Aspergillus species represents an important mycotic infection in public health concern as a human pathogen and as toxin-producing food contaminant. It releases a lot of spores which found in air, water, soil, plant debris, manure and animal feed. As fungal spore’s growing, it secretes digestive enzyme and mycotoxins leading to food spoilage and human mycotoxicosis [15].
Referring to the recorded results in table 3, Aspergillus sp. was prominently detected in breast samples other than wings and thighs samples, which came in agree with the previously recorded results of Darwish W, et al. (2016) [16] and Shaltout F, et al. (2019) [17] who found that the examined breast samples were more contaminated with fungal infection than wing and thigh samples. While the current prevalence of Aspergillus species in the examined samples came lower than those recorded by Hassan W, et al. (2019) [18] who found Aspergillus sp. in all the examined samples (100%) collected from Gharbiya governorate, Egypt. Moreover [19] also detected A. flavus and A. niger in 40 and 80% of the examined sausage samples of chicken origin, respectively
Referring to the obtained results of the microbiological identification of Aspergillus sp. isolates as recorded in table 4, they came in agree with the previously reported results by Darwish W, et al. (2016) [16] who found that A. niger was the predominant detected strain, followed by A. flavus and A. parasiticus in the examined samples of chicken cuts collected from Zagazig city, Egypt.
Some mold species can cause respiratory infections representing a significant risk for individual with severely weakened immune system [20]. Presence of mold in high incidence indicate bad hygienic measures adopted during handling, preparation and processing [21].
Mycotoxins are have been defined as naturally occurring secondary fungal metabolites produced in meat and meat products by direct growth of toxigenic molds such as Aspergillus species which produce aflatoxins and ochratoxins which threat public health due to their carcinogenic, hepatotoxic, nephrotoxic, teratogenic and mutagenic effects in humans and animals [22].
Aflatoxins are produced by a polyketide pathway that passes through about twenty-seven enzymatic reactions which have been regulated by sets of genes including nor-1, ver-1 and omtA have been shown to be involved in this process. aflD (nor-1) encodes a norsolorinic acid keto reductase needed for the conversion of the 10- keto group of Norsolorinic Acid (NOR) to the 10-hydroxyl group of Versicolorin A (VERA) [23] aflM (ver-1), predicted to encode a keto reductase, is involved in the conversion of VERA into Sterigmatocystin (ST) [24] ; aflP (omtA) codes for O-methyl transferase, which is one of the main genes responsible for transforming ST into O-methyl sterigmatocystin (OMST) that is the precursor for aflatoxin production [25].
Many other previous studies recorded the detection of these genes in their Aspergillus isolates of food origin with various PCR techniques [26-29], who conducted several studies investigating the aflatoxigenicity of Aspergillus sp. could detect different genes in their Aspergillus isolates.
Conclusion
It could be concluded that, breast samples revealed the highest contamination levels with Aspergillus sp; in addition, A. niger was the most prominently detected strain. PCR technique is a unique diagnostic tool for detection and identification of aflatoxigenic Aspergillus strains especially if the field of food safety. So, application of strict hygienic measures, proper use of water supply and food additives from good sources is recommended.
Article Information
Article Type: RESEARCH ARTICLE
Citation: Shaltout F, Heikal GI, Ghanem AM (2022) Molecular Detection of Aflatoxin Producing Aspergillus species Isolates in some Chicken Meat Cuts in Gharbiya Governorate, Egypt. Nutr Food Technol Open Access 8(1): dx.doi.org/10.16966/2470-6086.178
Copyright: © 2022 Shaltout F, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Publication history:
Received date: 01 Dec, 2021
Accepted date: 02 Feb, 2022
Published date: 14 Feb, 2022




