Introduction
The Flaviviridae family of icosahedral, enveloped (+) single stranded (ss) RNA viruses is responsible for serious diseases in human and other mammals. This family contains four genera, of which the largest, and most clinically relevant, Flavivirus, contains almost 70 members, most transmissible to humans by mosquitos or ticks. Among the mosquito- borne are the most virulent viruses like Dengue (DENV--the most serious threat), West Nile (WNV), Japanese encephalitis (JEV) and Yellow fever (YFV) [1].
Dengue is endemic in more than 100 countries with nearly 2/3 of the human population at risk and 40,000,000-reported cases/year (recent WHO statistics), incurring an estimate global healthcare burden of 9*109 USD per [2-5]. The four antigenically distinct serotypes (DENV 1-4) and multiple sub- phenotypes originated in different parts of the globe and they were mostly concentrated to the Asia-Pacific region [6]. However, in the last half century these serotypes have spread widely to endemic areas of the tropics and subtropics, thanks to an increase in global trade, transportation, human migration and urbanization, especially the emergence of densely populated quarters with low quality hygiene [7].
The linear, non-segmented dengue genome, a 10.7 kb (+ss) RNA, produces a 3391-amino acid long polyprotein that undergoes co- and post-translational processing by viral (NS2B-3) and cellular proteases to generate 3 structural proteins and 7 non-structural proteins [8- 11]. The mature virion contains the genetic material surrounded by the capsid protein and 180 copies of two glycoproteins. The amino terminal accounts for the structural proteins: Capsid (C), Membrane Precursor (prM) and Envelope (E), and the remaining genome gives rise to the non- structural proteins (NS1, 2A, 2B, 3, 4A, 4B, and 5) forming the viral Replication Complex (RC) [12,13].
Dengue 2 viruses can induce autophagy in several types of cells; this autophagy protects cells and increases viral replication [14]. The PERK pathway, an upstream effector of autophagy, mediates the induction of autophagy and protection [15]. Ataxia Telangiectasia Mutated (ATM) kinase, a DNA Damage Response protein, is activated early and may activate PERK and autophagy, leading to survival of infected cells [15,16]. The poorly studied non-structural protein 4A (NS4A) induces autophagy similarly to intact dengue virus and, like the intact virus; NS4A protects infected cells from toxins like Camptothecin (CPT), whereas other viral proteins do not [14]. NS4A-induced protection is mostly lost in cells that are deficient in autophagy, thus linking autophagy and flavivirus replication [14]. Macroautophagy, which recycles macromolecules and organelles [17,18], is most active in a stressed, starved or an infected cell and includes: 1) Nucleation of phagophore, (recruitment of membrane); 2) Expansion of the phagophore membrane around the target ‘cargo’ by LC3II and ATG9 proteins; 3) Closure of the membrane into the autophagosome; 4) Fusion of the autophagosome and lysosome into an autolysosome and 5) Degradation of the cargo and permease-mediated efflux into cytosol [19-21]. The prime mediators of autophagy signaling are sensors of cell energy (AMPK-Adenosine Monophosphate-Activated Protein Kinase) and nutrition (mTOR-Mammalian Target of Rapamycin). In a healthy cell, mTOR phosphorylates ULK1 and ATG13, blocking autophagy, whereas the upstream AMPK removes mTOR suppression, inducing autophagy [22-25]. Ataxia Telangiectasia Mutated (ATM), a nuclear protein, upregulates AMPK and thus autophagy [26,27]. ATM is induced by DNA damage, reactive oxygen and nitrogen species (ROS and RNS), and hypoxia and exported from nucleus, blocking the activity of mTOR through a phosphorylation cascade involving AMPK and other enzymes [20,24,26,28-34].
The virus can persist and be shed in seemingly recovered patients or mosquito cells, providing an important means by which epidemics are initiated and sustained [35-38]. The protection of infected cells by virus provides a means by which the virus can persist. Here we measure the persistence of the different serotypes in MDCK cells. Dengue virus 1-4 can persist in cultured cells for as much as two weeks, and the viruses and their NS4A proteins, though differing in amino acid sequence, can all induce autophagy and protect renal epithelial cells from stressors that would otherwise induce apoptosis. Thus autophagy is a target pathway to design antiviral therapeutics.
The objective of this study is to identify pro-survival mechanisms common to all DENV serotypes and their NS4A proteins. Since we have linked induction of autophagy to increased viral infection in our previous studies [14,15], we looked at the relation between autophagy and its upstream regulators in cells infected with these serotypes. NS4A was identified as the sole contributor towards DENV-induced autophagy in our previous study [14], hence we also tested the ability of NS4As from all these serotypes to induce autophagy and protect cells from toxins. Our results provide with an important clue for development of antiviral therapy targeted towards all the serotypes.
Experimental Procedures
Cell culture and treatment
MDCK (gift of Dr. Anastasia Gregoriades, Queens College, Flushing, NY), HeLa: GFP-LC3 (provided by Guido Kroemer, Institut Gustave-Roussy, Villejuif, France), were maintained at 37°C in Dulbecco’s minimum essential media (DMEM) with 10% FBS, 1% penicillin/streptomycin. C6/36 mosquito cells (Adolfo Garcia-Sastre, Mount Sinai Medical School, New York, who also provided Dengue virus and pCAGGS-HA plasmid vector) were maintained at 28°C in Eagle’s minimal essential media Sigma), 1% nonessential amino acids (Sigma), 2 mM L-glutamine and 25 units/ml fungizone (Invitrogen), and 50 units/ml penicillin/streptomycin), both in 5% CO2 in air. ATM inhibitor caffeine (Sigma) was used at a final concentration of 20 µM, which does not provoke autophagy [39]. Camptothecin (CPT, Sigma) was used at a final concentration of 5 µM. Cells were incubated with inhibitors for 1 h prior to infection.
Viral infection
Cells were seeded and allowed to attach overnight in maintenance media and washed with 1X phosphate-buffered saline (PBS) before infecting at an MOI (Multiplicity Of Infection)=5 unless otherwise stated. Mock-infection involved treatment with virus media lacking virus then addition with new media. Virus stocks were diluted with ice-cold flavivirus diluting media (1X PBS containing 0.75% bovine serum albumin fraction V, pH 8.0). Following application of dilutions of virus, cells were incubated for 2 hours. Cells were then covered with maintenance media, and incubated until data collection.
Propagation of virus
For expansion of stocks of dengue-2 virus, sub-confluent C6/36 mosquito cells were infected with virus and incubated for 6 days. The culture medium was agitated and collected, then mixed 2:1 with flavivirus freeze media (0.75% bovine serum albumin fraction V in 0.12 M NaCl, 0.05 M H3 BO3, pH 9.0), and stored at -80°C. Viral load was then determined by plaque assay [40]. Plaques were counted and virus titer was determined. Each sample was run in triplicate.
Transfection of mammalian cells
NS4A from MODOC and DENV 1-4 were expressed in mammalian (HeLa, MDCK) cells using a pCAGGS-HA plasmid vector. TOPOplasmid vectors were used as negative controls. Cells were plated to 85% confluence and allowed to attach overnight in penicillin/ streptomycin-free cell type-appropriate culture media supplemented with 10% FBS. Cells were transfected using the Lipofectamine 3000 system (Thermo Fisher Scientific) according to the manufacturer’s protocol. Post-transfection, cells were incubated at 37°C for at least 16 h before collection or treatment.
Bioinformatics analysis of dengue and Modoc NS4A sequences
Dengue NS4A sequences were obtained from GenBank (GenBank ID): dengue 1 (AHI43749.1), dengue 2 (AHI43753.1), dengue 3 (AHL17464.1) and dengue 4 (AHN50410.1). Modoc NS4A (NP_740265.2) was obtained from NCBI. NS4A protein sequence alignments were carried out according to Mathew, et al. [41] and Trojnar, et al. [42] using the Geneious Basic 5.0.3 software. Phylogenetic trees were constructed using Jukes Cantor, UPGMA tree build method. Bootstrap analysis of phylogenetic trees was performed with 1000 trials and 111 random seeds..
Assessment of cell viability
Cells were infected and exposed to CPT at 24 HPI (Hours PostInfection), incubated for an additional 24 h at 37ºC, 5% CO2 , collected by trypsin digestion, and stained with 0.4% trypan blue in 1X PBS [43]. Live (white) and dead (blue) cells were counted on a hemocytometer with cell viability expressed as [(% dead cells)-(% dead in controls)].
Western blot
At 48 HPI, cells were scraped and washed with ice-cold 1X PBS before whole lysate proteins were collected in RIPA buffer and quantified using Bio-Rad protein assay and an Ultrospec III spectrophotometer (GE Healthcare). Western blots were performed by SDS-PAGE [43], using β tubulin (Sigma-Aldrich) as loading control and primary antibody for LC3B (Sigma-Aldrich). Positive signals were detected using ECL (GE Healthcare or Cell Signaling) and visualized using hyper film ECL photo radiographic film (GE Healthcare).
Immunocytochemistry
Cells were seeded onto flame-sterilized glass coverslips (Globe Scientific Inc.), allowed to attach overnight, and infected and treated as described above. After the treatment/infection, cells were washed with 1X PBS and fixed with fresh, ice-cold 4% Paraformaldehyde (Fisher) for 10 minutes, and washed with 1X PBS again. The coverslips were then mounted on glass slides (Globe Scientific Inc.), using Fluoromount (Sigma). Embedded cells were then observed by confocal (Leica, Germany) or fluorescence (Leitz) microscopy.
Results
Different serotypes of dengue virus protect renal kidney cells and induce autophagy. Of the four well-studied serotypes of dengue, we previously established that dengue 2 induces autophagy, which protects cells from death induced by other toxins or stressors [14]. The protection conferred by autophagy is most likely the mechanism allowing cells to continue to multiply and produce dengue virus. We asked if the induction of autophagy and protection was unique to serotype 2 or whether it was characteristic of this group of flavivirus. MDCK cells were infected with dengue 1, 2, 3 or 4 at MOI=5 for 24 h then challenged with 5 µM CPT for another 24 hours. It must be noted that when MDCK were infected with dengue 2 and monitored for infectivity for up to 3 weeks, the infected cells still produced mature virus at a good titer (data not shown). We also observed similar persistent infectivity with other serotypes such as DENV3 (data not shown). As shown in figure 1A, none of the serotypes of virus killed significantly more cells than the mock-infected controls (≤ 10% death after 48 HPI) as measured by trypan blue assay. On the other hand, after 24 h, CPT kills approximately 45% of the MDCK cells. However, infection of cells with any single dengue serotype reduces CPTinduced cell death to amounts similar to treatment with virus alone (+ Den1, 2, 3, 4). Thus all 4 serotypes protect MDCK cells from death induced by CPT.
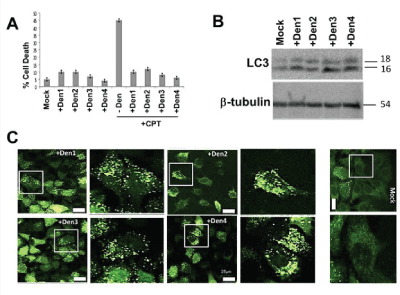
Figure 1: The serotypes 1-4 of dengue virus protect renal kidney cells against camptothecin and increase autophagy in them. (A) Trypan blue exclusion assay shows high viability in CPT-exposed MDCK cells infected with dengue 1-4 (MOI 5, 48 HPI), reducing killing within 24 h from 45% to 5-15%, equivalent to that of infection alone. (B) Western blot of whole cell lysates from mock-infected and DENV-infected MDCK shows increased lipidation (16 kDa LC3-II) in MDCK cells infected with any serotype compared to mock-infection (Mock). β-tubulin was used as a loading control. (C) HeLa cells stably expressing GFP-LC3 (HeLa:GFP-LC3) show increased GFP-LC3 puncta formation after infection with any single serotype (+Den), consistent with increased endogenous LC3-II, indicating increased autophagy. Right column is a 3X (linear) enlarged portion of picture on the left, indicated by box.
To determine whether the protection conferred by infection with any dengue serotype also derived from upregulation of autophagy, as previously observed for dengue 2 [14], we examined by western blot the amount of endogenous LC3-II. Whole cell lysates from uninfected and mock-infected cells show low endogenous LC3-II in mock-infected cells after 48 HPI, reflecting basal level (housekeeping) autophagy (Figure 1B). However, infection with each individual strain of dengue virus increases LC3-II compared to mock-infected samples (Figure 1B). The induction of autophagy by each serotype was further confirmed using HeLa cells that stably express GFP-LC3 (HeLa: GFPLC3). In this system translocation of LC3-II to, and accumulation on, autophagosome membranes result in a punctate distribution of GFP. Mock-infected HeLa cells show predominantly diffused GFPLC3 with few LC3-GFP puncta, indicating low autophagy typical of healthy cells (Figure 1C). As was the case for dengue 2 viruses, dengue serotypes 1, 3 and 4 increase formation of puncta, suggesting induction of autophagy (Figure 1C), a result that is congruent with the results from western blot (Figure 1B). We conclude that up- regulation of autophagy and protection of MDCK cells are conserved among the several dengue strains.
NS4As of Dengue virus 1-4 upregulate autophagy
NS4A is a small hydrophobic protein previously shown to be the only flavivirus gene capable of inducing autophagy, which in turn protects against toxins like Camptothecin (CPT), Staurosporine (STS), Cycloheximide (CHX) and even biological inducers of apoptosis like influenza A virus [14].
To determine whether the function of NS4A (induction of autophagy/ cell protection) is shared among the different dengue serotypes, we used the Geneious Basic 5.0.3 software program to compare the sequence homology among the NS4As of different serotypes. Amino acid homology of NS4A among dengue serotypes is moderate (43%), and higher than between dengue and Modoc (compare Figures 2A and 2B). Phylogenetic analysis of dengue 1-4 NS4A shows a close relationship among the dengue serotypes compared to Modoc (Figure 2B), consistent with the accepted phylogeny of these members of Flaviviridae. NS4As from all four dengue serotypes are more closely related to each other, than they are to the Modoc NS4A. This has been shown by the 100% consensus support for grouping together all dengue NS4A and low substitution per site within dengue serotypes (Figure 2B, consensus support=node values, substitution per site=horizontal values). Of the dengue strains, dengue 1 and 3 are sister taxa that were geographically distinct strains forming a monophyletic clade, all stemming from a single recent common ancestor [44]. Dengue 1 and 3 NS4A are related at 88.6% consensus support while dengue 2 and 4 are clustered together within the four strains at 75.8% consensus support. The poor homology of dengue to Modoc virus resembles those in other distantly related Flaviviruses including hepatitis C, Japanese encephalitis and West Nile virus, which often serve as out-group [14,45].
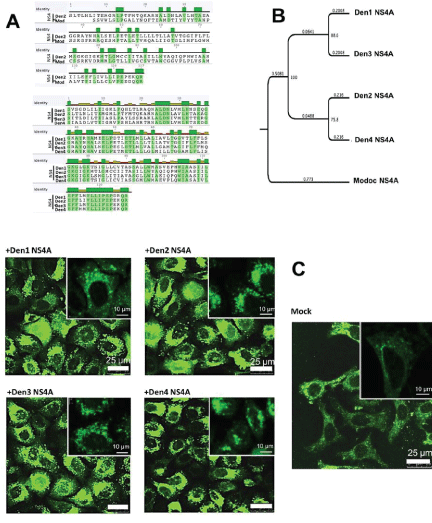
Figure 2: The NS4A proteins of dengue virus 1-4 are moderately conserved. (A) Dengue virus 2 and Modoc NS4A proteins, both known to induce autophagy, show 21% similarity indicating poor conservation (green=conserved amino acids). Among dengue specifically, 43% of amino acids are shared among the NS4As of different serotypes. (B) The NS4As of dengue virus serotypes 1-4 are more closely related to each other than to NS4A of Modoc. (C) Transient expression of NS4As from all dengue serotypes in HeLa:GFP-LC3 cells produces more GFPLC3 puncta formation and more puncta-positive cells compared to mock-infected cells. Scale bars indicate 25 µm for the larger photographs and 10 µm for the insets.
We examined induction of autophagy by LC3 translocation in HeLa:GFP-LC3 after transfection with plasmids containing NS4A genes from dengue 1,2,3 or 4. Control mock-transfected (plasmid vector alone) cells display housekeeping LC3-GFP puncta (Figure 2C, Mock). All the dengue virus serotypes NS4As induce LC3-GFP puncta formation compared to control, in a large subset of transfected cells, indicating an upregulation in autophagosome formation (Figure 2C + Den1-4 NS4A). These results, like those above, indicate that the ability of all serotypes of NS4A to induce autophagy in renal and HeLa cells is conserved. In spite of their differences, NS4A of all the DENV serotypes as well as Modoc induce autophagy and protect cells against CPT-induced death, to nearly similar degrees [14,46].
In MDCK cells, ATM kinase is important for NS4A-induced autophagy and protection from CPT
Active ATM kinase increases autophagy and protects cells infected with whole dengue 2 virus [15]. DEN2 and MODOC NS4A both protect MDCK cells against CPT, correlating with a modest increase in autophagy [15]. We tested whether ATM kinase would do the same in cells in which NS4A was over expressed as opposed to infecting the cells. NS4A gene from either DENV2 or Modoc (murine flavivirus) was transfected into MDCK cells that had been pretreated with the ATM inhibitor caffeine, with or without the toxin CPT. In presence of the NS4A gene, MDCK cells were protected from the toxic effect of CPT; however, this protection by NS4A was partially eliminated by caffeine. Cell death in CPT-treated and NS4A-transfected cells almost doubled in the presence of caffeine, compared to CPT-treated and NS4A-transfected cells that were not treated with caffeine (Figure 3A).
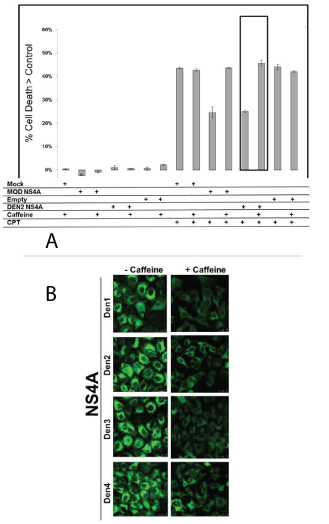
Figure 3: In MDCK cells, ATM kinase is needed for NS4A-induced autophagy and protects against CPT. (A) MDCK cells were transfected with plasmids coding for MODOC and DEN2 NS4A before being exposed to CPT, caffeine or both CPT and caffeine. Over expression of the viral gene protected cells from CPT. However, blocking ATM kinase (by caffeine) diminished the protection. [Mod=Modoc NS4A, Empty=empty vector] (B) HeLa:GFP-LC3 cells were pretreated with caffeine before being transfected with the NS4A gene from all dengue serotypes. As in figure 2C, NS4A over expression induces autophagy. However, caffeine suppresses the number of GFP-LC3 puncta.
We next evaluated autophagy. We tested whether NS4A protein, like intact DENV2, could mediate ATM-dependent autophagy in HeLa: GFP-LC3 cells. We pretreated HeLa: GFP-LC3 cells with caffeine before transfecting them with NS4A-containing plasmids which, as shown in figure 2C, induce autophagy as measured by an increase in the count of GFP-LC3 puncta. Pretreatment with caffeine sharply decreases the number of GFP-LC3 puncta. Thus, like the whole virus, all NS4As induce autophagy (Figure 3B) and protect CPT-treated cells, a response that is partially overcome by blocking ATM kinase with caffeine.
Discussion and Conclusion
Flaviviruses are important for human health and, with the recent emergence of Zika and the likelihood that it can persist in asymptomatic individuals, we need to determine the mechanisms by which Flaviviruses survive, replicate, and persist in mammalian cells. These viruses replicate in double-membrane compartments like the endoplasmic reticulum and autophagosomes. We suggest that they use the autophagic pathway to persist in cultured cells and, potentially, in individuals. Thus the importance of autophagy to the survival and persistence of the virus needs to be resolved.
Autophagy is important to the pathogenicity of dengue virus. DENV2 infection activates ATG5-dependent autophagy in hepatic (Huh7) and fibroblast (MEF) cells leading to increased titer of virus. Blocking autophagy in Huh7.a.1, BHK21 cell lines and AG129 mice resulted in a heat-sensitive and non-infectious dengue virion. Deficiency of autophagy related (ATG) genes or specific autophagy inhibitors like spautin-1 reduce dengue replication, leading to temperature sensitive, mutant virions, and reducing survival of infected cells [47-49]. However, the relationship between autophagy and DENV infection depends on the infected cell type. In human monocytes (U937), which the virus does not directly infect, induction of autophagy decreased viral replication. DENV2-mediated autophagy protects canine kidney epithelial (MDCK) and mouse fibroblast (MEF) cells from toxic stimuli but does not protect murine macrophages [14], in which infection leads to apoptotic cell death. Thus it is important to look at the effects of infection on the status of autophagy in cells from different sources [47-49].
Dengue serotypes use the NS4A proteins to induce autophagy, and at least one serotype (DENV2) achieves this by controlling signals for both upregulation (ATM, ER stress and ROS production) and down regulation (mTOR) of autophagy [14,15]. This increase in autophagy augments resistance to toxic insults by limiting activation of apoptosis and promotes further replication of virus. Since PI3K-dependent autophagy correlates with increased cell survival, the identification of NS4A as the major factor helps to define the pathways that are relevant to pathology of Dengue virus. Here we establish that NS4As from all four serotypes induce autophagy by activating ATM, which possibly upregulates the AMPK-LKB1 axis [50]. Pharmacological activation of the AMPK protein reduces the infectivity of three Flaviviruses, DENV, WNV and ZIKV, indicating that the anti-viral aspect of this protein is conserved among Flaviviruses [51]. Thus, more mechanistic studies should be conducted on the relationship among DDR, ER stress (UPR/PERK) and the pro-survival homeostatic processes (autophagy) in DENV-infected cells. On the clinical front, persistence of virus in seemingly recovered individuals creates a reservoir for the accidental infection of contacts and for the emergence of future epidemics. Large numbers of mature virions persist in infected canine kidney cells 3-4 weeks after infection (data not shown). The ability of the virus to persist in vitro provides a means of studying persistence in vivo. Persistence varies by cell type and might depend on ATM-mediated autophagy. Since autophagy is responsible for increased cell protection, this might serve as a useful model to study the mechanisms behind persistent infection.
Finally, the presence of homology (43%) and strong phylogenetic relationship among the NS4As from different dengue serotypes, suggests a possible target domain that induces autophagy and protection in MDCK cells, which might be identified to limit the activity of NS4A. Thus our study offers an alternative approach to vaccine design, and presents autophagy as a target pathway to design antiviral therapeutics. This common feature of all serotypes may prove to be a target for polyvalent therapies.
Our findings are also important because we identify a common pathway (ATM-induced autophagy) that is conserved not only among different DENV serotypes but also their NS4A proteins. This result might prove useful to design a pan-DENV therapy where hurdles like Antibody-Dependent Enhancement (ADE) can be bypassed. Most vaccines are not effective against secondary infection because of ADE, where antibodies developed during primary infection facilitate secondary infection. Since we have identified specific signaling pathways responsible for viral persistence and infectivity, our findings could offer a viable alternative to traditional vaccine-Antiviral therapy aimed at metabolic targets utilized by the virus.
The current study paves the way for more investigation of signaling pathways conserved among cells infected by different serotypes. One link that needs to be validated is that between autophagy flux and viral infectivity. Although we have established the role of ATM kinase in inducing autophagy and protecting cells, whether the autophagy flux is essential for viral production or whether it is merely a byproduct of viral infection must be verified. Future studies should be oriented toward gaining a deeper understanding of the effect of autophagymediated cell protection on viral infectivity.