Abstract
Ribosome profiling (RP) is a technique of sequencing of mRNA regions occupied by ribosomes at a given time. This recently invented technique has been well appreciated with regard to our deeper understanding of the phenomenon of codon usage bias in organisms. Along with the information about gene expression, which is studied by the commonly used techniques such as Microarray and RNA sequencing, RP can give information about ribosome speed along a coding sequence. The technique has helped scientists to identify many non-canonical translational initiation sites, pausing events along a coding sequence and stop codon read-through that contribute to the gene expression at the level of translation. In this review, RP methodology, outcomes and software packages available for data analysis are discussed.
Keywords
RNA sequencing; Ribosome; Coding; RP methodology
Introduction
The central dogma of molecular biology states that the genetic information contained within DNA is transcribed to messenger RNA (mRNA), which is translated into polypeptide chain and folded into functional proteins [1]. Unlike DNA inside the cell, transcriptome varies according to the requirement of the cell. Therefore, the study of gene expression by analyzing entire transcripts can provide important insights into the spatial and temporal gene regulation inside a cell. Microarray and RNA sequencing (RNA-seq) are two extensively used techniques to analyze genome wide abundances of transcripts. It is pertinent to note that regulation of gene expression occurs both at transcriptional level as well as post transcriptional/translational levels. The latter one can`t be determined by the two above-mentioned techniques. So, to distinguish which region on mRNA molecules are engaged in active translation, recently a novel technique has been developed, known as Ribosome profiling (RP). RP is defined as a genome wide mapping of nucleotides on mRNA occupied by ribosomes during translation at a specific time. Generally polysome profiling is used to distinguish mRNA molecules involved in active translation from the non-translated mRNA. But polysome profiling is unable to answer several questions such as (a)whether ribosome speed is uniform along the mRNA during translation or across different mRNA molecules?,(b)is there a connection between speed of ribosome and gene expression?,(c) how mRNA structures affect ribosome speed during translation? And (d) how ribosome speed during translation is influenced by the initiation and termination codons? However, the RP technique can address the abovementioned queries in a genome wide context. In table 1 the advantages of RP technique over Microarray, polysome profiling and RNA sequencing have been mentioned in brief.
| Microarray |
RNA-seq |
Polysome profiling |
RP |
| Profiling of known genes using pre synthesized probes |
Profiling whole Transcriptome |
It can assay mRNA that is actively involved in translation by looking at polysomes |
Profiling translating transcriptome |
| Analyze mRNA expression based on nucleic acid hybridization and fluorescence |
Analyze mRNA expression by next generation sequencing |
Identification and expression profiling of mRNA under translation by RNA-seq |
Identification and quantification of regions of mRNA bound to ribosomes by next generation sequencing |
| Identification of novel transcripts is not possible |
Can identify novel transcripts, alternative splicing and SNPs |
Can identify novel transcripts and alternative splicing, SNPs |
Can identify novel transcripts, SNPs |
| Provide information of transcription level |
Provide information of transcription level |
Provides post transcriptional information but unable to decipher location of ribosomes along mRNA molecules |
Provides post transcriptional information and positional information of ribosomes along mRNA molecules |
Table 1: Distinctive advantages of RP techniques over other currently used techniques such as Microarray, RNA-seq, Polysome profiling
RP Method
RP is a method that can decode the finer level of translational regulation in which the same transcriptome can produce distinctive translating transcriptomes. Introduced by Ingolia et al. [2], it is a robust technique to sequence ribosome protected mRNA fragments known as “ribosome footprints”, using high throughput deep sequencing technology (Figure 1). In RP experiment, cells are initially treated with translational inhibitor e.g. cycloheximide to freeze ribosomes as is where basis to capture snapshot of translational status under a given condition. The fixed cells are then lysed, digested with nonspecific nuclease such as RNase or micrococcal nuclease (MNase) to degrade unprotected mRNA fragments and leaving intact ribosome occupied fragments of a size of approximately 28-30 nucleotides. The derived small fragments are processed with the deep sequencing protocol to obtain ribosome footprints. RP is a highly sensitive technology that can even detect rare translational messages in position specific manner by analyzing ribosome footprint density along mRNA molecules. Mapping of these footprints to genome wide ribosome occupancy profiles eventually decode the ribosome positions at nucleotide level. Besides its recent origin, this technology is widely accepted and adopted to profile ribosome footprints of diverse organisms in various physiological conditions [3]. Several optimization of the original protocol by Ingolia et al. [2] have been reported recently [4- 7]. Till date, only Illumina (California, USA) provides commercial kits with brand name TruSeqRibo Profile for RP in mammalian and yeast system. One of the difficulties of the technique is to avoid contamination from the rRNA. Effective control of rRNA fragments in RP experiments is essential, because the background noise due rRNA fragments can reduce the efficiency of detecting the signals from mRNA. Therefore the commercially available kit like Ribo-Zero rRNA depletion kit (Epicenter) is generally used for removal of rRNA contaminating fragments in RP experiments. Recently, double strand specific nuclease was used for rRNA depletion in different species [8].
RP Gives Insight in to Different Levels of Translation
Ribosomes binding at ribosome binding site (RBS) of mRNA and incorporation of first amino acid mark the start of translational initiation. Initiating ribosomal machinery continues translation until it reaches the stop codon, i.e. elongation and finally nascent polypeptides released by release factor to terminate translation. Now, it is evident that RP provide new information’s regarding translational dynamics at three different levels i.e. Initiation, elongation and termination and that is discussed below.
Mapping of Translational Initiation Sites
Mapping of translational start site or initiation site (TIS) on mRNA, which defines the open reading frame (ORF) in a gene was not easy until the RP technology came into existence. Measuring or mapping translational initiation sites. For mapping the translational initiation sites harringtonine and lactimidomycin drugs were used to block ribosomes initiating translation or otherwise premature termination of translation can be achieved by puromycin followed by blocking with cycloheximide treatment [9]. Global translational initiation sequencing (GTI-seq) uses E site inhibitor drugs cycloheximide (CHX) and lactimidomycin (LTM), to identify noble TIS in a transcriptome by analyzing ribosome footprints in mammalian cells [10]. As quantification of TIS is not possible with GTI-seq, a new technique was adopted by Gao et al. [11], called quantitative translation initiation sequencing (QTI-seq), which uses LTM to freeze initiating ribosomes followed by puromycin (PMY) treatment for depleting elongating ribosomes to quantify and map TIS. From the recent RP experiments primarily targeted to map TIS, it becomes very clear that the mechanism of translational initiation is far more complex than that was previously thought. Dramatic increase in the number of non-AUG start or initiation at 5ʹ UTR sites are documented through RP experiments [12-14] and these non-AUG translational initiation mostly found in short upstream ORFs (uORF). Short uORF are found in the upstream region of genes, which undergo translation but unable to produce functional protein product because of very small size, instead they play regulatory role by inhibiting the translation of the downstream protein coding gene or the main ORF [15]. Non upstream regulatory ORFs (nORF) have initiation signals inside the main ORF also documented from RP experiments. Analysis of ribosome footprint data, revealed that translation of functionally inactive novel ORFs from long interspersed non coding RNA (lincRNA) [16]. In bacteria, Tetracycline mediated inhibition of ribosomes by blocking ribosomes at initiation sites effectively maps several new TIS using RP [17].
Ribosome Dynamics During Elongation
By RP technique, the ribosome occupying regions during translation elongation are mapped. Therefore mRNA regions where ribosome remains for a longer duration during translation are represented more in ribosome profiling in comparison to the mRNA regions where ribosome remains for a lesser duration. The former regions represents regions where the translation speed is slower (otherwise known as ribosome pausing site) which may be due to the occurrence of non-optimal codons or due to mRNA secondary structures or due to anti Shine-Dalgarno sequence or interaction of nascent peptide with the ribosome, presence of stretches containing amino acid such as proline, serine and histidine [18,19] etc Whereas the latter regions represents where the translation speed is higher due to the presence of the optimal codons or absence the above hindering factors on mRNA or growing polypeptide. Ribosome pausing is important for co-translational protein folding [20].
Differential usage of synonymous codons in an organism is known as codon usage bias (CUB) [21]. By comparing synonymous codon usage between the high expression genes and the low expression genes within a genome, scientists have predicted that certain synonymous codons are decoded faster than the other synonymous codons during translation [22]. This discrepancy among the synonymous codons with regard to translation speed got experimental evidence recently by RP [23-25]. conducted an experiment using codon manipulated luciferase mRNA fragments in Neurosporacrassa cell free systems and concluded that corelation exists between codon usage and translational dynamics i.e. optimal codon increase the translational elongation speed, while non-optimal codon reduces the rate of translational elongation (Figure 2). Non optimal codon induced ribosome pausing can result in co-translational folding of the derived protein which is an important mechanism to maintain proper structure and function of proteins [26]. In an earlier study, codon optimization in N.crassa circadian clock gene frequency (frq), found that optimization in disordered regions can result in structurally different and nonfunctional protein product [25,27].
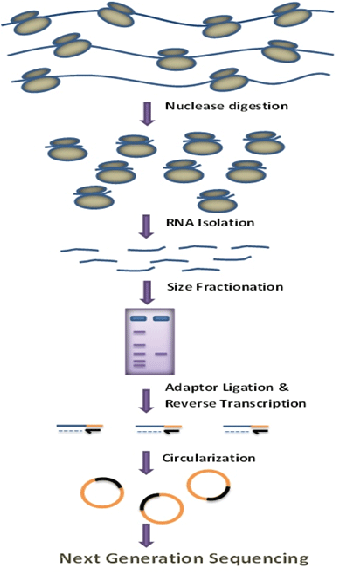
Figure 1: Schematic representation of RP method
Different steps of Ribosme profiling are as follows: (i) isolation of ribosome bound mRNA from cell; (ii). digestion by RNase such that ribosome bound RNA remains protected and enzyme accessible RNA regions are digested; (iii). Isolation of the ribosome bound RNA and synthesis of cDNA followed by ligation with adapter and next gen sequencing.
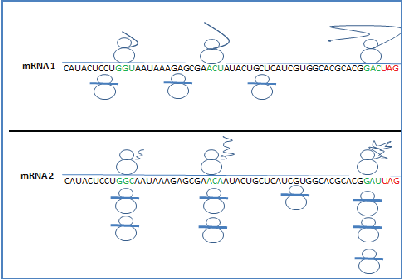
Figure 2: Schematic representation of protein synthesis and ribosome footprints of two mRNA molecules i.e. mRNA1 with all optimal codons and mRNA2 with both optimal and non-optimal codons demarcated by a black line. In both cases, upper part showing translating ribosomes with synthesized protein, lower part showing ribosome footprint density. In mRNA1, presence of optimal codons throughout the mRNA, induces fast translation. In mRNA2, due to presence of synonymous mutations in some codons induces ribosome pausing and co-translational folding. Synonymous mutations resulting in non-optimal codons in the mRNA showed more ribosome footprints and ribosome halts in those positions for cognate tRNA molecules.
Termination of Translation and Stop Codon Read-Through
Stop codons contain termination signals for any given protein synthesis event. But in some instances these stop codons doesn’t terminate the protein synthesis, as a consequence, a portion or the whole 3 ʹun-translated region is also getting translated to produce protein variants with extended C-terminal region. Stop codon read-through is known to occur in some viruses [28,29]. RP technique assists to decipher many new stop codon read-through events in yeast, Drosophila and mammals [30-32]. Stopcodon read-through as a result of ribosomal frame shifts was reported in ribosome footprints in 3 ʹends of transcripts of Drosophila melanogaster [31]. Stop codon read-through imparts altered functions and localization of the subsequent protein product and thereby also plays important role in evolution of species.
Determination of A and P Sites in Footprints
Ribosome footprints are generally ~28 nucleotides in length but some footprints are not of canonical length. Due to footprint length heterogeneity [33-35], it is difficult to design a method or software to predict the position of acceptor (A) and peptidyl (P) sites within a footprint. Earlier, A site determined either by aligning 5’end of footprint with initiation sites of transcripts or central nucleotides of a footprint considered as A site [9]. A recent study by Martens et al. [36], ascertained the A and P site locations in ribosome footprints based on paused sites at specific amino acid such as serine, histidine, proline etc. In the study, separate location of A and P sites within footprints revealed for eukaryotes and prokaryotes. P and A site were predicted 4 and 5 codons downstream from 5’ end in yeast whereas 5 and 4 codon upstream from the 3’ end in E. coli. From the abovementioned study, it is evident that pausing tendency at certain codons can give indication of A and P sites within a ribosome footprint.
Ribosome Profiling of Organelle
Organelle such as mitochondria and chloroplast gene expression also constitutes integral share for viability of organisms. These organelles have their own protein synthesis machinery, which is more similar to prokaryotes than eukaryotes. As organellar ribosomes are different from cytoplasmic ribosomes, so conventional ribosome profiling may skip organellar Ribosome Protected Fragments. Rooijers et al. [37] successfully optimized protocol for mitochondrial RP to profile wild type and disease triggering mitochondrial translation. Mitochondrial RP was also carried out for understanding the dynamics of yeast mitochondrial biogenesis [38]. Similarly, chloroplast RP was carried out at different stages of chloroplast differentiation in maize by using ribosome profiling [39]. With some technical improvements in experimental procedure and bioinformatics analysis RP technique can be very useful for understanding of translation dynamics in mitochondria and chloroplast.
Software Packages for Analysis of RP Data
RP is accepted by molecular biologists as a promising technology to unfold translational regulation. The analysis of RP data resulting from RNA-seq platform which consists of large amount of short read data, is still a daunting task for a biologist. With increasing popularity of RP technology several software packages and pipelines were developed for quantitative and qualitative analysis of data generated from RP experiments [41-44]. Though these pipelines minimize many inherent problems due to very short reads but footprints derived from similar transcripts with different 5’ and 3’ UTR regions can’t be mapped with 100% confidence level. Ribo Taper is a recently developed pipeline which can easily and effectively predict protein coding ORF using triplet periodicity. Ribo Tools is an open source quality monitoring toolbox available from Galaxy, developed for minimizing uncertainty by precisely determining codon occupancy profiles, translation ambiguities and stop codon read through in RP data [42]. Several other pipelines for analysis of RP data are also mentioned in the table 2.
Software packages |
Script |
Availability |
Released under |
Reference |
| RiboTaper |
R |
https://ohlerlab.mdc-berlin.de/ software/RiboTaper_126/ |
Github |
Calviello et al. [3] |
| riboseqR |
R |
http://bioconductor.org/packages/release/bioc/ html/riboSeqR.html |
Bioconductor |
Chung et al. [8] |
| RiboTools |
Python 2.7 |
https://testtoolshed.g2.bx.psu.edu/view/ rlegendre/ribo_tools |
Galaxy |
Legendre et al. [42] |
| PROTEOFORMER |
Pearl5 |
http://www. biobix.be/proteoformer |
Galaxy |
Crappe et al. [41] |
| GWIPS-viz |
|
http://gwips.ucc.ie/ |
UCSC genome browser |
Michel et al. [9] |
Table 2: Software packages and pipelines available for analysis of RP data
Conclusion
RP adopts RNAseq technologies to understand complexities or dynamics associated with translational initiation, elongation and termination. Transcriptome sequencing using RNA seq, in parallel with RP is a preferred method for understanding transcriptional and translational dynamics of a given organism in a particular condition. In near future we will get more insight into the codon-anticodon interaction difference by ribosomes of different organisms. Availability of commercially available kits with simpler workflows and automated pipeline for analysis will definitely help scientist envisioned to profile diverse organisms and will enhance our understanding of translational dynamics.
Article Information
Article Type: Review Article
Citation: Agarwala N, Ray SK (2016) Ribosome Profiling: An Insight into the Dynamics of Translating Ribosomes on a Coding Sequence: A Review Article. Int J Mol Genet Gene Ther 2(1): doi http://dx.doi.org/10.16966/2471-4968.107
Copyright: © 2016 Agarwala N, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Publication history:
Received date: 30 Sep 2016
Accepted date: 25 Oct 2016
Published date: 31 Oct 2016



