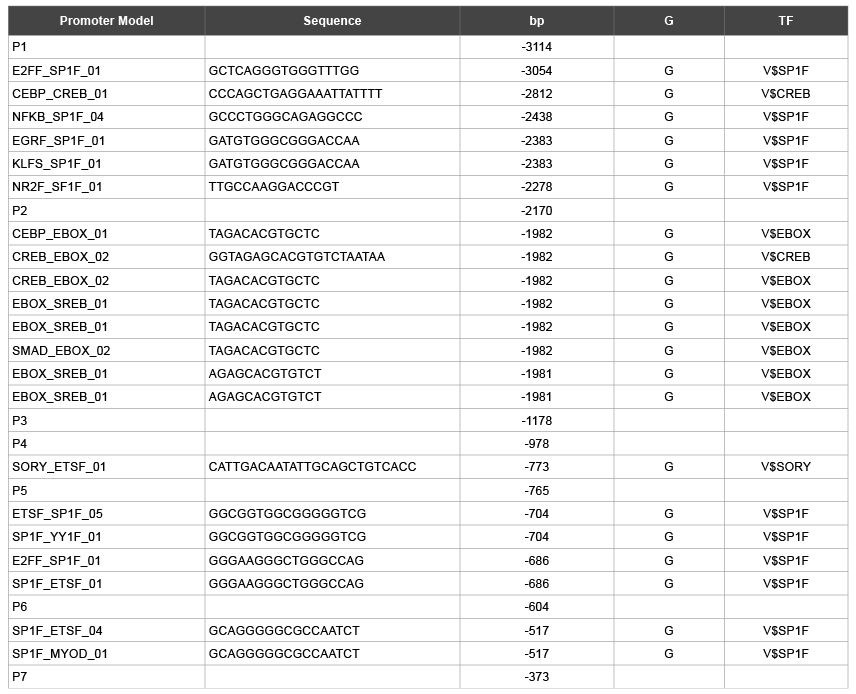
Table 1: Glycosylation site on promoter model of mMC3r

Xin WEN1,2* Wen Wei ZENG3
1Developmental Neurobiology Section, NHLBI, NIH, Bethesda, MD 20892, USA*Corresponding author: Xin WEN, Developmental Neurobiology Section, NHLBI, NIH, UGO, Eunice Kennedy Shriver, NICHD, NIH, Bethesda, MD 20892, USA, Tel: +1 240 506 3095; E-mail: xin.wen@nih.gov
Objective: This study is to understanding the transcription profile of nociception-related melanocortin receptor MC1R, obesity related neural melanocortin receptor Mc3r and MC4R.
Methods: For MC1R, Mc3r and MC4R, relative luciferase activity RLA and the distribution of phosphorylation [P], glycosylation sites [G] on transcription factor for promoter model were plotted on the same chart.
Results: Within the 3.2 kb upstream of the methionine ATG, for all three receptors, trend lines of RLA with that of [G] or ([P]-[G]), both tend to have negative reciprocal relationship. It brings up the question: for the neural receptors MC1R, Mc3r and MC4R, does the nutrition and obesity related glycosylation regulated the transcriptional activity in a negative reciprocal way?
Conclusion: The negative reciprocal relation between trend line of [G] or ([P]-[G]) and that of RLA, give a specific digit evidence for the first time to the theory, which the structure related glycosylation and the signal sensing phosphorylation, exhibit either independently or Interactively on regulation of transcription activity.
Luciferase activity; Melanocortin receptor; Transcription factor
RLA: Relative luciferase activity; DLA: Dual luciferase assay; MC1R: Human melanocortin 1 receptor; Mc3r: Mouse melanocortin 3 receptor; MC4R: Human melanocortin 4 receptor; Promoter model: A promoter model represents a framework of two or more conserved elements (e.g., transcription factor binding sites) with a defined distance (and strand orientation); TF: Transcription factor; P1: construct Plasmid 1; P7: construct Plasmid 7; P9: construct Plasmid 9; MC1R-P1: construct Plasmid 1 for human melancortin 1 receptor; MC1R-P9: construct Plasmid 9 for human melancortin 1 receptor; Mc3r-P1: construct Plasmid 1 for mouse melanocortin 3 receptor; Mc3r-P7: construct Plasmid 7 for mouse melanocortin 3 receptor; MC4R-P1: construct Plasmid 1 for human melanocortin 4 receptor; MC4R-P5: construct Plasmid 5 for human melanocortin 4 receptor; [P]: number of phosphorylation sites on TF for promoter models; [G]: number of glycosylation sites on TF for promoter models; [P]-[G]: number of difference between phosphorylation and glycosylation sites on TF for promoter models; [P]+[G]: number for sum of phosphoylation and glycosylation sites on TF for promoter models
MC1R may be involved in the inflammatory aspects of pain signaling [1]. In the central nervous system (CNS), MC1R mRNA expression and protein levels are detected in the hypothalamus, amygdala, locus coeruleus and dorsal raphe nucleus, which are areas of the brain that are implicated in regulating emotion and stress. Therefore, MC1R not only functions in skin pigmentation, nociception and anti-inflammatory response, but also in melanocortin pathway involving motivation, reward and emotional states, and mood disorders such as anxiety and depression.
And Mc3r [2] is widely distributed in the Central Nerve System; it is involved in controlling ingestive behaviors and autonomic function. The central and peripheral Mc3rS are involved in the maintaining normal body weight.
Also, MC4R are distributed [1] in cortex, hypothalamus, brainstem, spinal cord, astrocytes, heart and lungs. Mutations of the MC4R gene that lead to reduced receptor functioning have been consistently associated with obesity. Obesity has been linked to MC4R signaling, suggesting that stimulation of this receptor promotes compulsive grooming and its blockade reverses the induction of this excessive behavior in rodents.
In addition, Mc3r and MC4R are in central melanocortin system [3,4], which involved in regulation of thermogenesis, body weight regulation through its role in appetite and energy expenditure, and dysregulation of the energy homeostasis, contributing to the pathogenesis of diseases such as cardiovascular, cerebrovascular diseases and pathophysiological process of cachexia, as well as to the type 2 diabetes and obesity.
On the other hand, primary gene induction or repression in eukaryotes does not require protein synthesis [5], suggesting the involvement of posttranslational modifications [6,7]. Since many different types of stimuli that affect gene expression also lead to the activation of protein kinases, analysis of transcription factor phosphorylation is essential for complete understanding of the signal pathways. The activity of transcription factors may be modulated by their signal-sensing domain including phosphorylation [8]. In addition, as nutrient sensitive sugar modification, glycosylation, interfere with the epigenetic control of gene expression.
MC1R, Mc3r, and MC4R receptors are involved in anxiety, depression, cardiovascular and cachexia diseases as well as obesity, mutations and polymorphisms of melanocortin-1 receptor type 1 gene (MCR1) play a role in skin cancer [9]. Moreover, phosphorylation or glycosylation are perhaps required for the activation of transcription factors. However, the regulatory mechanism and post translation modification in the epigenetic transcription regulation of these Melanocortin Receptors has yet to be fully investigated.
For Mc3r, determination of transcription starting site of Mc3r using 5’RACE was performed on total Mus musculus hypothalamus RNA using the 5’RLM-RACE kit [10] (nature protocol exchange), Series deletion fragments containing sequences extending from 3084 bp to 283 bp were subcloned into the upstream of the firefly luciferase gene in pGL3 vector, as confirmed by restriction enzyme digestion and sequencing.
And study of transcriptional activation and regulation of Mc3r was performed by using gene reporter assay [5], human HEK293 was transiently transfected using FuGENE’s 6 Tranfection Reagent, 48 hours after transfection and over 65% cell confluency, Renila-Firefly Dual luciferase activity was measured. Assay was tested by triplicates on independent experiments. The activities of all constructs in pGL3 basic in HEK293 were measured. Afterwards, Promoter model [11] [A promoter model represents a framework of two or more conserved elements (e.g., transcription factor binding sites) with a defined distance (and strand orientation). Usually, promoter models are much more specific than single elements like transcription factor binding sites. Therefore, a promoter model can give higher evidence that the matching sites are functional] was inspected by Genomatix (http://www.genomatix. com/). The glycosylation (Table 1) and phosphorylation sites (Table 2) on the promoter model were searched by Protein Knowledge bases of Uniprot (http://www.uniprot.org/).
For MC1R, Relative Luciferase Activity (in SK-Mel-2 human melanoma cells) is from figure 1 in Osamu Moro’s paper [12], and the sequence information for the corresponding constructs is from figure 2 in the same paper. Glycosylation (Table 3) and phosphorylation sites (Table 4) of promoter models were searched in the same way as for that of Mc3r.
For MC4R, Relative Luciferase Activity of HEK 293 is from figure 1 of Christian Vaisse’s paper [13], and DNA sequence is from figure 3 in the same paper. Glycosylation (Table 5) and phosphorylation sites (Table 6) of promoter models were searched in the same way as for that of Mc3r and MC4R.
For MC1R, as it shown in figure 2a, MC1R-P1 has RLA of 100.0, with length of 2918 bp, ranging from -3200 bp to -282 bp, there are 25 phosphorylation sites and 11 glysosylation sites on TF for promoter model between MC1R-P1 and MC1R-P2. MC1R-P2 has RLA of 130.0, with length of 2131 bp, ranging from -2413 bp to -282 bp; there are 33 phosphorylation sites and 8 glysosylation sites on TF for promoter model between MC1R-P2 and MC1R-P3. MC1R-P3 has RLA of 94.0, with length of 1100 bp, ranging from -1382 bp to -282 bp; there are 30 phosphorylation sites and 7 glysosylation sites on TF for promoter model between MC1R-P3 and MC1R-P4. MC1R-P4 has RLA of 131.0, with length of 648 bp, ranging from -930 bp to -282bp; there are 19 phosphorylation sites and 2 glysosylation sites on TF for promoter model between MC1R-P4 and MC1R-P5. MC1R-P5 has RLA of 116.0, with length of 284 bp, ranging from -566 bp to -282 bp; there are 2 phosphorylation sites and 1 glysosylation site on TF for promoter model between MC1R-P5 and MC1R-P6. MC1R-P6 has RLA of 88.4, with length of 235 bp, ranging from -517 bp to -282 bp; there are 5 phosphorylation sites and 3 glysosylation sites on TF for promoter model between MC1R-P6 and MC1R-P7. MC1R-P7 has RLA of 22.0, with length of 177 bp, ranging from -459 bp to -282 bp; there is no phosphorylation site and no glysosylation site on TF for promoter model between MC1R-P7 and MC1R-P8. MC1R-P8 has RLA of 3.0, with length of 165 bp, ranging from -447 bp to -282 bp; there is no phosphorylation site and no glysosylation site on TF for promoter model between MC1R-P8 and MC1R-P9. MC1R-P9 has RLA of 3.6, with length of 132 bp, ranging from -414 bp to -282bp; there are 22 phosphorylation sites and 6 glysosylation sites on TF for promoter model on MC1R-P9.
For MC1R, If plot the RLA data and {the glycosylation site [G], phosphorylation site [P], sum of [P]+[G], difference of [P]-[G] site} of TF for promoter model on the same chart, got figure 3, the polynomial behavior of MC1R is: trend line of RLA is (y=-2E-10x4 - 1E-06x3 -0.002x2 -2.313x-546.1); trend line of [G] is (y=1E-11x4 +8E- 08x3 +0.000x2 +0.120x+29.89); trend line of [P]-[G] is (y=3E-11x4 +2E- 07x3 +0.000x2 +0.272x+63.35); trend line of [P] is (y=5E-11x4< +3E- 07x3 +0.000x2 +0.393x+93.25); trend line of [P]+[G] is (y=6E-11x4 +4E- 07x3 +0.000x2 +0.513x+123.1). Trend line RLA and trend line of ([G], [P], [P] ± [G]) give an inverted image to each other. In detail, the sign of leading coefficient is opposite to each other, and sign of coefficient of corresponding degree are also opposite to each other. Furthermore, trend line of RLA (y=-2E-10x4 -1E-06x3 -0.002x2 -2.313x-546.1) and trend line of [G] (y=1E-11x4 +8E-08x3 +0.000x2 +0.120x+29.89) tend to have a relation of negative reciprocal (Figure 3b). At the same time, trend line of RLA (y=-2E-10x4 -1E-06x3 -0.002x2 -2.313x-546.1) and trend line of [P]-[G] (y=3E-11x4 +2E-07x3 +0.000x2 +0.272x+63.35) tend to have a relation of negative reciprocal as well (Figure 3a).
For Mc3r, as it indicated in figure 2b, Mc3r-P1 has RLA of 0.964, with length of 2998 bp, ranging from -3114 bp to -116 bp, there are 29 phosphorylation sites and 6 glysosylation sites on TF for promoter model between Mc3r-P1 and Mc3r-P2. Mc3r-P2 has RLA of 1.342, with length of 2054 bp, ranging from -2170 bp to -116 bp; there are 15 phosphorylation sites and 8 glysosylation sites on TF for promoter model between Mc3r-P2 and Mc3r-P3. Mc3r-P3 has RLA of 2.596, with length of 1062 bp, ranging from -1178 bp to -116 bp; there are 4 phosphorylation sites and no glysosylation site on TF for promoter model between Mc3r-P3 and Mc3r-P4. Mc3r-P4 has RLA of 2.844, with length of 862 bp, ranging from -978 bp to -116 bp; there are 11 phosphorylation sites and 1 glysosylation site on TF for promoter model between Mc3r-P4 and Mc3r-P5. Mc3r-P5 has RLA of 3.010, with length of 649 bp, ranging from -765 bp to -116 bp; there are 13 phosphorylation sites and 4 glysosylation sites on TF for promoter model between Mc3r-P5 and Mc3r-P6. Mc3r-P6 has RLA of 3.098, with length of 488 bp, ranging from -604 bp to -116 bp; there are 8 phosphorylation sites and 2 glysosylation sites on TF for promoter model between Mc3r-P6 and Mc3r-P7. Mc3r-P7 has RLA of 1.845, with length of 257 bp, ranging from -373 bp to -116 bp; there are 10 phosphorylation sites and no glysosylation site on TF for promoter model on Mc3r-P7.
For Mc3r, If plot the RLA data and {the glycosylation site [G], phosphorylation site [P], sum of [P]+[G], difference of [P]-[G] site} of TF for promoter model on the same chart, got figure 1, the polynomial behavior of Mc3r is: trend line of RLA is (y=-2E-11x4 -6E-08x3 -6E- 05x2 -0.033x-3.146); trend line of [P]+[G] is (y=1E-13x5 +7E-10x4 +1E- 06x3 +0.001x2 +0.404x+55.37); trend line of [G] is (y=2E-14x5 +1E-10x4 +2E- 07x3 +0.000x2 +0.033x+1.873); trend line of [P] is (y=9E-14x5 +5E-10x4 +1E- 06x3 +0.001x2 +0.371x+53.50); trend line of [P]-[G] is (y=7E-14x5 +4E- 10x4 +9E-07x3 +0.000x2 +0.337x+51.63). Trend line RLA and trend line of ([G], [P], [P] ± [G]) give an inverted image to each other. In detail, the sign of leading coefficient is opposite to each other, and sign of coefficient of corresponding degree are also opposite to each other. Furthermore, trend line of RLA (y=-2E-11x4 -6E-08x3 -6E-05x2 -0.033x-3.146) and trend line of [G] (y=2E-14x5 +1E-10x4 +2E-07x3 +0.000x2 +0.033x+1.873) tend to have a relation of negative reciprocal (Figure 1b). At the same time, trend line of RLA (y=-2E-11x4 -6E-08x3 -6E-05x2 -0.033x-3.146) and trend line of [P]-[G] (y=7E-14x5 +4E-10x4 +9E-07x3 +0.000x2 +0.337x+51.63) tend to have a relation of negative reciprocal as well (Figure 1a).

Table 1: Glycosylation site on promoter model of mMC3r
For MC4R, as shown in figure 2c, MC4R-P1 has RLA of 14.7, with length of 2510 bp, ranging from -2926 bp to -416 bp; there are 12 phosphorylation sites and no glysosylation site on TF for promoter model between MC4R-P1 and MC4R-P2. MC4R-P2 has RLA of 19.8, with length of 1510 bp, ranging from -1926 bp to -416 bp; there are 14 phosphorylation sites and 1 glysosylation site on TF for promoter model between MC4R-P2 and MC4R-P3. MC4R-P3 has RLA of 22.3, with length of 760 bp, ranging from -1176 bp to -416 bp; there are 7 phosphorylation sites and 1 glysosylation site on TF for promoter model between MC4R-P3 and MC4R-P4. MC4R-P4 has RLA of 20.8, with length of 140 bp, ranging from -556 bp to -416 bp; there are 4 phosphorylation sites and 1 glysosylation site on TF for promoter model between MC4R-P4 and MC4R-P5. MC4R-P5 has RLA of 9.0, with length of 60 bp, ranging from -476 bp to -416 bp; there are 10 phosphorylation sites and 2 glysosylation sites on TF for promoter model on MC4R-P5.
For MC4R, If plot the RLA data and {the glycosylation site [G], phosphorylation site [P], sum of [P]+[G], difference of [P]-[G] site} of TF for promoter model on the same chart, got figure 4, the polynomial behavior of MC4R is: trend line of RLA is (y=-2E-10x4 -9E-07x3 -0.001x2 -1.234x-279.0); trend line of [G] is (y=3E-11x4 +1E-07x3 +0.000x2 +0.117x+28.39); trend line of [P]-[G] is (y=3E-14x5 +3E-10x4 +9E-07x3 +0.001x2 +0.753x+169.0); trend line of [P] is (y=4E-14x5 +3E-10x4 +1E-06x3 +0.001x2 +0.870x+197.4); trend line of [P]+[G] is (y=4E-14x5 +3E-10x4 +1E-06x3 +0.001x2 +0.988x+225.8). Trend line RLA and trend line of ([G], [P], [P] ± [G]) give an inverted image to each other. In detail, the sign of leading coefficient is opposite to each other, and sign of coefficient of corresponding degree are also opposite to each other. Furthermore, trend line of RLA (y=-2E-10x4 - 9E-07x3 -0.001x2 -1.234x-279.0) and trend line of [G] (y=3E-11x4 +1E- 07x3 +0.000x2 +0.117x+28.39) tend to have a relation of negative reciprocal (Figure 4b). At the same time, trend line of RLA (y=-2E-10x4 -9E-07x3 - 0.001x2 -1.234x-279.0) and trend line of [P]-[G] (y=3E-14x5 +3E-10x4 +9E- 07x3 +0.001x2 +0.753x+169.0) tend to have a relation of negative reciprocal as well (Figure 4a).
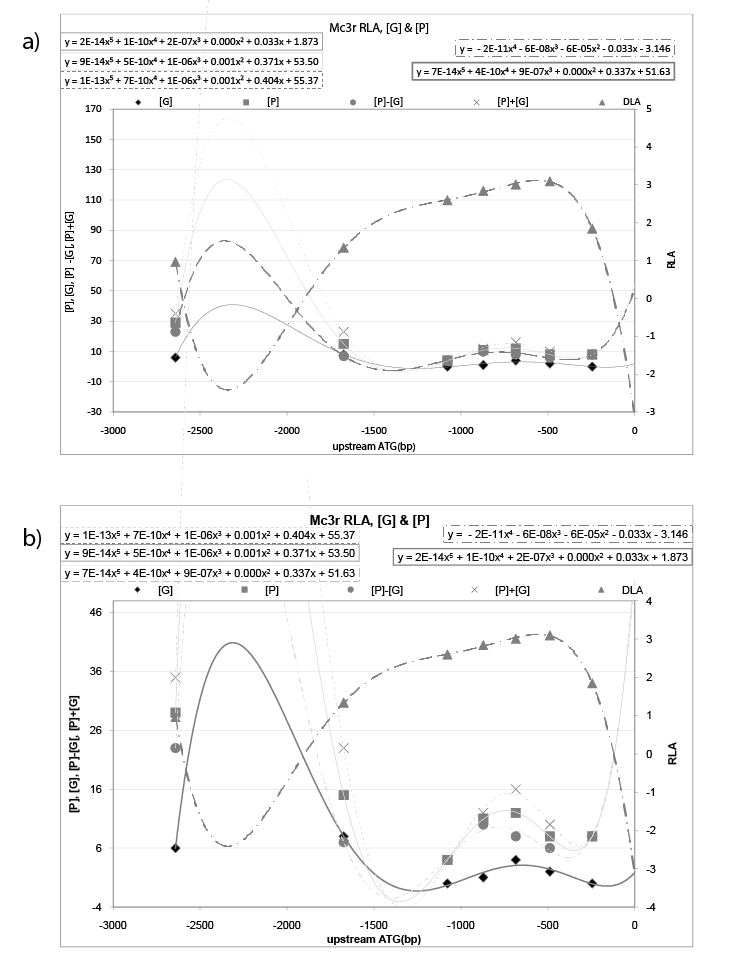
Figure 1: RLA data of Mc3r and {the glycosylation site [G], phosphorylation site [P], sum of [P]+[G], difference of [P]-[G] site} of TF for promoter model. For Mc3r, plot the RLA data and {the glycosylation site [G], phosphorylation site [P], sum of [P]+[G], difference of [P]-[G] site} of TF for promoter model on the same chart, the polynomial behavior of Mc3r is: trend line of RLA is (y=-2E-11x4 -6E-08x3 -6E-05x2 -0.033x-3.146); trend line of [P]+[G] is (y=1E-13x5 +7E-10x4 +1E-06x3 +0.001x2 +0.404x+55.37); trend line of [G] is (y=2E-14x5 +1E-10x4 +2E-07x3 +0.000x2 +0.033x+1.873); trend line of [P] is (y=9E-14x5 +5E-10x4 +1E-06x3 +0.001x2 +0.371x+53.50); trend line of [P]-[G] is (y=7E-14x5 +4E-10x4 +9E-07x3 +0.000x2 +0.337x+51.63). Trend line RLA and trend line of ([G], [P], [P] ± [G]) give an inverted image to each other. In detail, the sign of leading coefficient is opposite to each other, and sign of coefficient of corresponding degree are also opposite to each other. Furthermore, trend line of RLA and trend line of [P]-[G] tend to have a relation of negative reciprocal (a). Concurrently, trend line of RLA and trend line of [G] tend to have a relation of negative reciprocal as well (b).
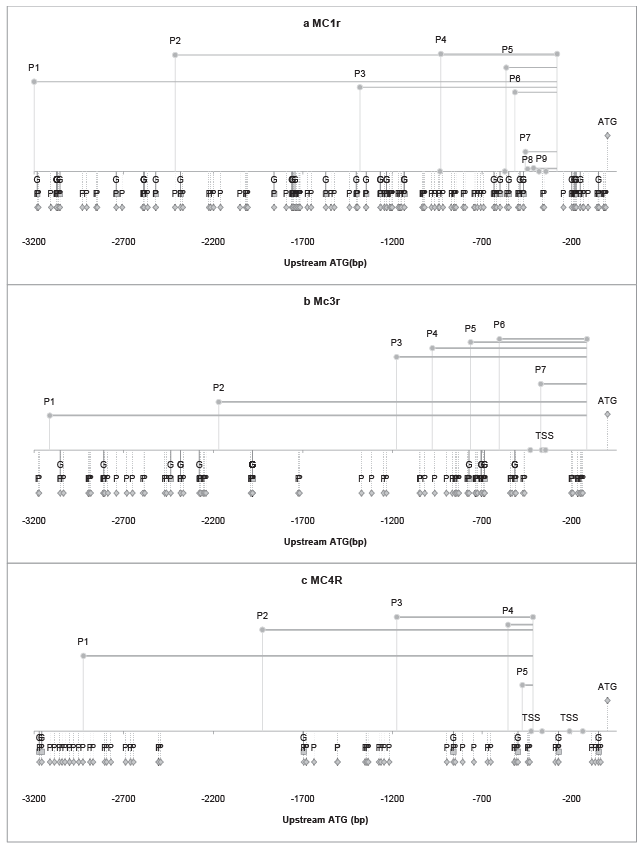
Figure 2: RLA of corresponding plasmids for (a) MC1R (b) Mc3r (c) MC4R and distribution of phosphorylation site and glycosylation site on TF for promoter model. The RLA of the corresponding plasmids is plotted on upper Y axis (Y>0) with the corresponding sequence of sense primer and that of antisense primer, concurrently, a horizontal line is drawn to link the starting point and ending point of each constructed plasmid respectively. On the other hand, within the 3200 base pair (bp) upstream the methionine-ATG (0 bp), on the lower Y axis area (Y<0), the phosphorylation (P) sites and glycosylation (G) sites is also pinned on the corresponding sequence (bp) for transcription factors promoter models. It shows the distribution of P and G. And Transcription Starting Site (TSS) is plotted on X axis (Y=0). (a) MC1R, plasmids (P1, P2, P3, P4, P5, P6, P7, P8, P9). (b) Mc3r, plasmids (P1, P2, P3, P4, P5, P6, P7). (c) MC4R, plasmids (P1, P2, P3, P4, P5).
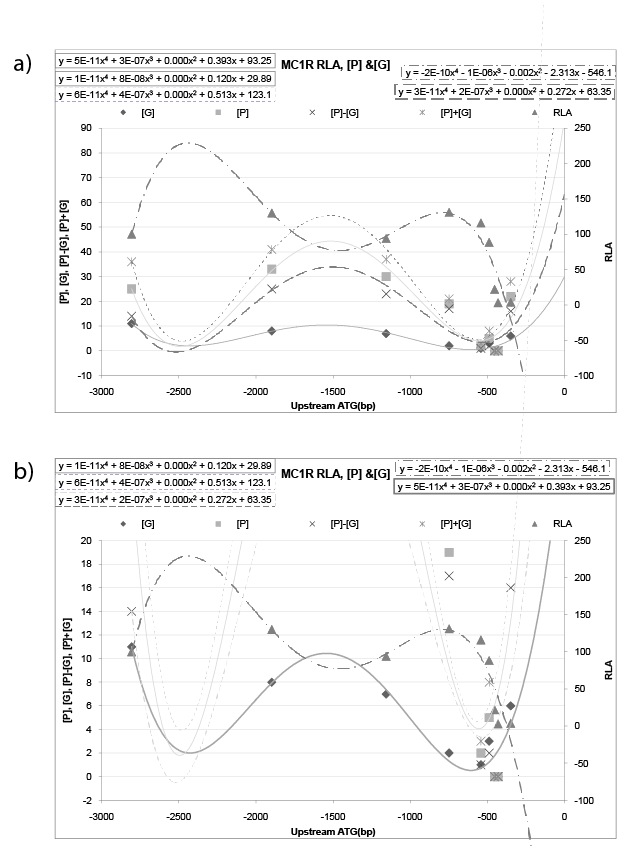
Figure 3: RLA data of MC1R and {the glycosylation site [G], phosphorylation site [P], sum of [P]+[G], difference of [P]-[G] site} of TF for promoter model. For MC1R, plot the RLA data and {the glycosylation site [G], phosphorylation site [P], sum of [P]+[G], difference of [P]-[G] site} of TF for promoter model on the same chart, the polynomial behavior of is: trend line of RLA is (y=-2E-10x4 -1E-06x3 -0.002x2 -2.313x-546.1); trend line of [G] is (y=1E- 11x4 +8E-08x3 +0.000x2 +0.120x+29.89); trend line of [P]-[G] is (y=3E-11x4 +2E-07x3 +0.000x2 +0.272x+63.35); trend line of [P] is (y=5E-11x4 +3E- 07x3 +0.000x2 +0.393x+93.25); trend line of [P]+[G] is (y=6E-11x4 +4E-07x3 +0.000x2 +0.513x+123.1). Trend line RLA and trend line of ([G], [P], [P] ± [G]) give an inverted image to each other. In detail, the sign of leading coefficient is opposite to each other, and sign of coefficient of corresponding degree are also opposite to each other. Furthermore, trend line of RLA (y=-2E-10x4 -1E-06x3 -0.002x2 -2.313x-546.1) and trend line of [P]-[G] (y=3E- 11x4 +2E-07x3 +0.000x2 +0.272x+63.35) tend to have a relation of negative reciprocal (a). Concurrently, trend line of RLA (y=-2E-10x4 -1E-06x3 -0.002x2 - 2.313x-546.1) and trend line of [G] (y=1E-11x4 +8E-08x3 +0.000x2 +0.120x+29.89) tend to have a relation of negative reciprocal as well (b).
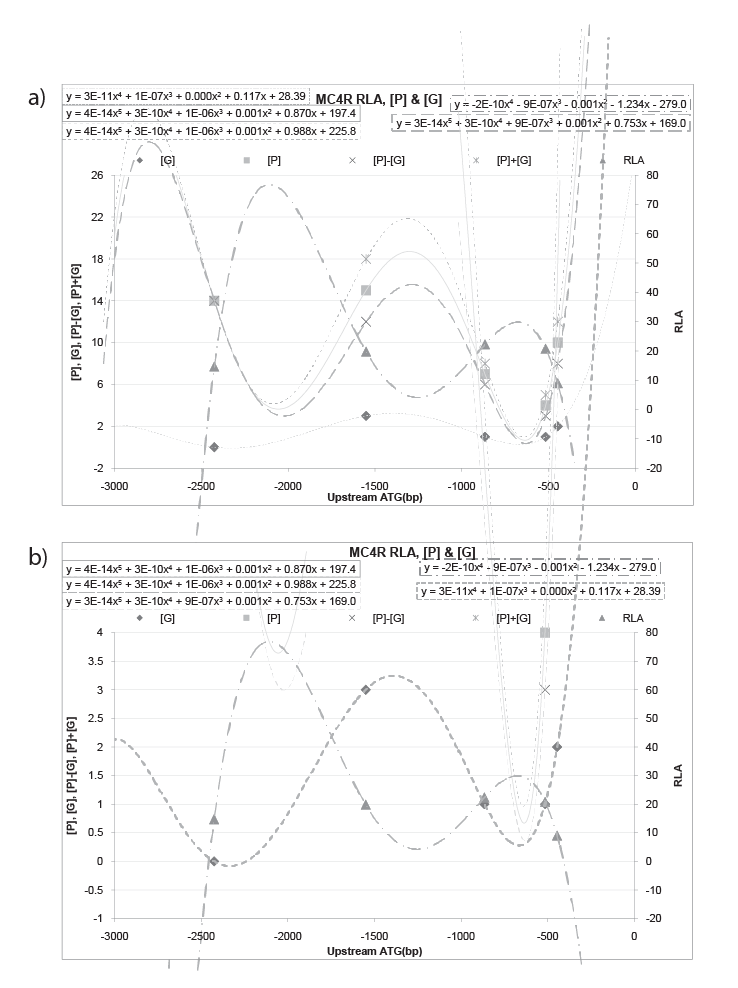
Figure 4: RLA data of MC4R and {the glycosylation site [G], phosphorylation site [P], sum of [P]+[G], difference of [P]-[G] site} of TF for promoter model. For MC4R, plot the RLA data and {the glycosylation site [G], phosphorylation site [P], sum of [P]+[G], difference of [P]-[G] site} of TF for promoter model on the same chart, the polynomial behavior of MC4R is: trend line of RLA is (y=-2E-10x4 -9E-07x3 -0.001x2 -1.234x-279.0); trend line of [G] is (y=3E-11x4 +1E-07x3 +0.000x2 +0.117x+28.39 ); trend line of [P]-[G] is (y=3E-14x5 +3E-10x4 +9E-07x3 +0.001x2 +0.753x+169.0); trend line of [P] is (y=4E-14x5 +3E-10x4 +1E-06x3 +0.001x2 +0.870x+197.4); trend line of [P]+[G] is (y=4E-14x5 +3E-10x4 +1E-06x3 +0.001x2 +0.988x+225.8). Trend line RLA and trend line of ([G], [P], [P] ± [G]) give an inverted image to each other. In detail, the sign of leading coefficient is opposite to each other, and sign of coefficient of corresponding degree are also opposite to each other. Furthermore, trend line of RLA and trend line of [P]-[G] tend to have a relation of negative reciprocal (a). Concurrently, trend line of RLA and trend line of [G] tend to have a relation of negative reciprocal as well (b).
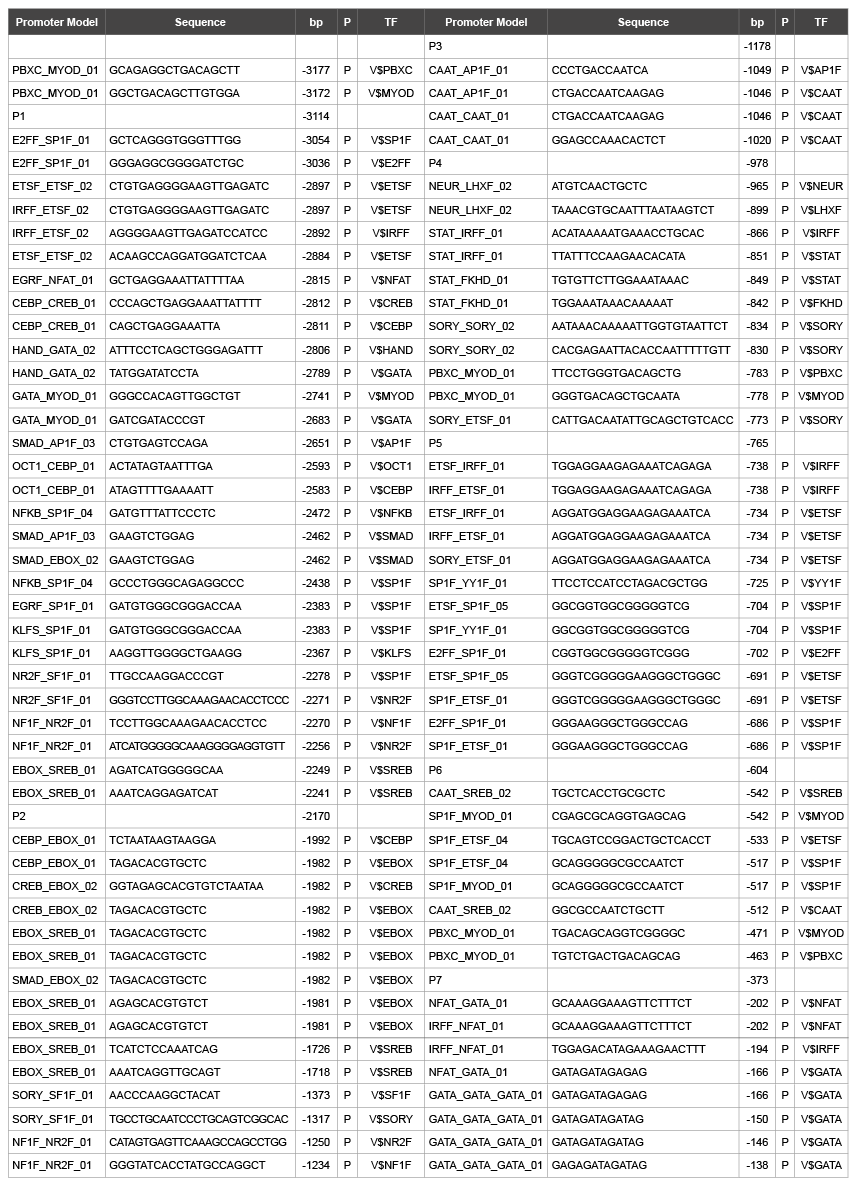
Table 2: Phosphorylation site on promoter model of mMC3r
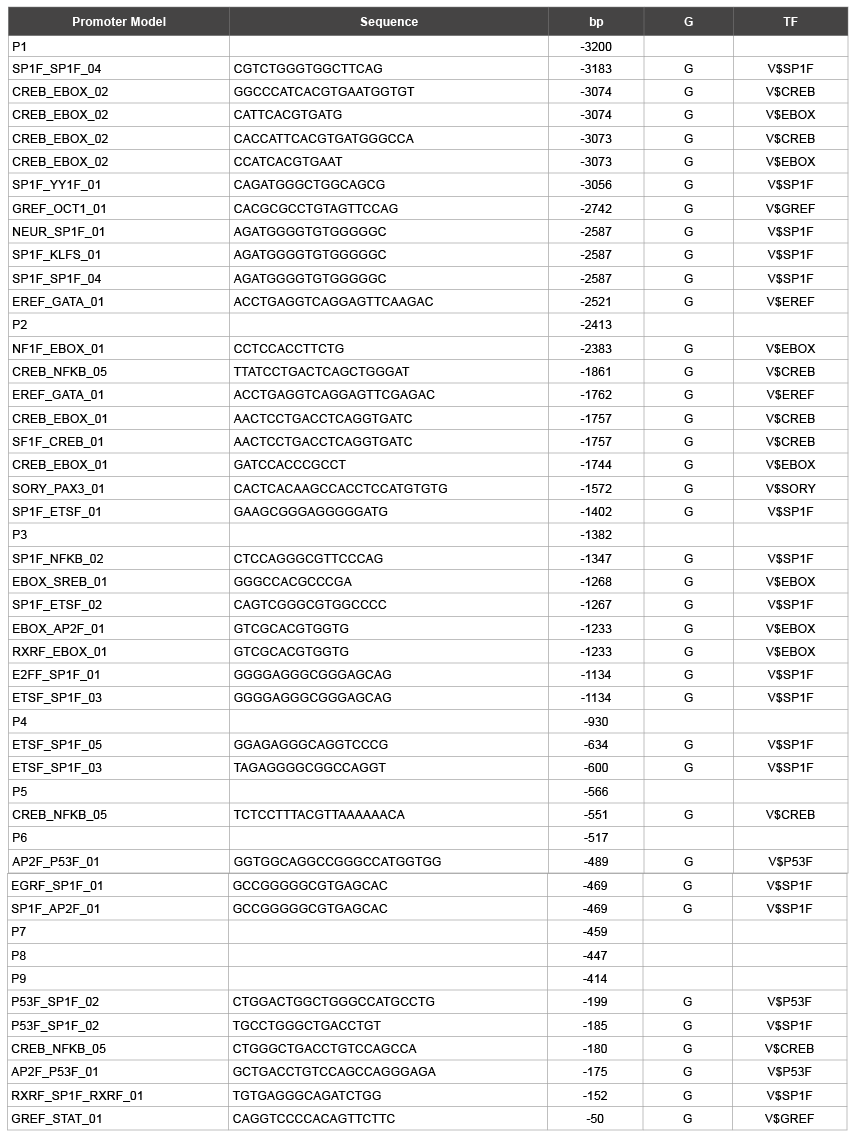
Table 3: Glycosylation site on promoter model of hMC1R
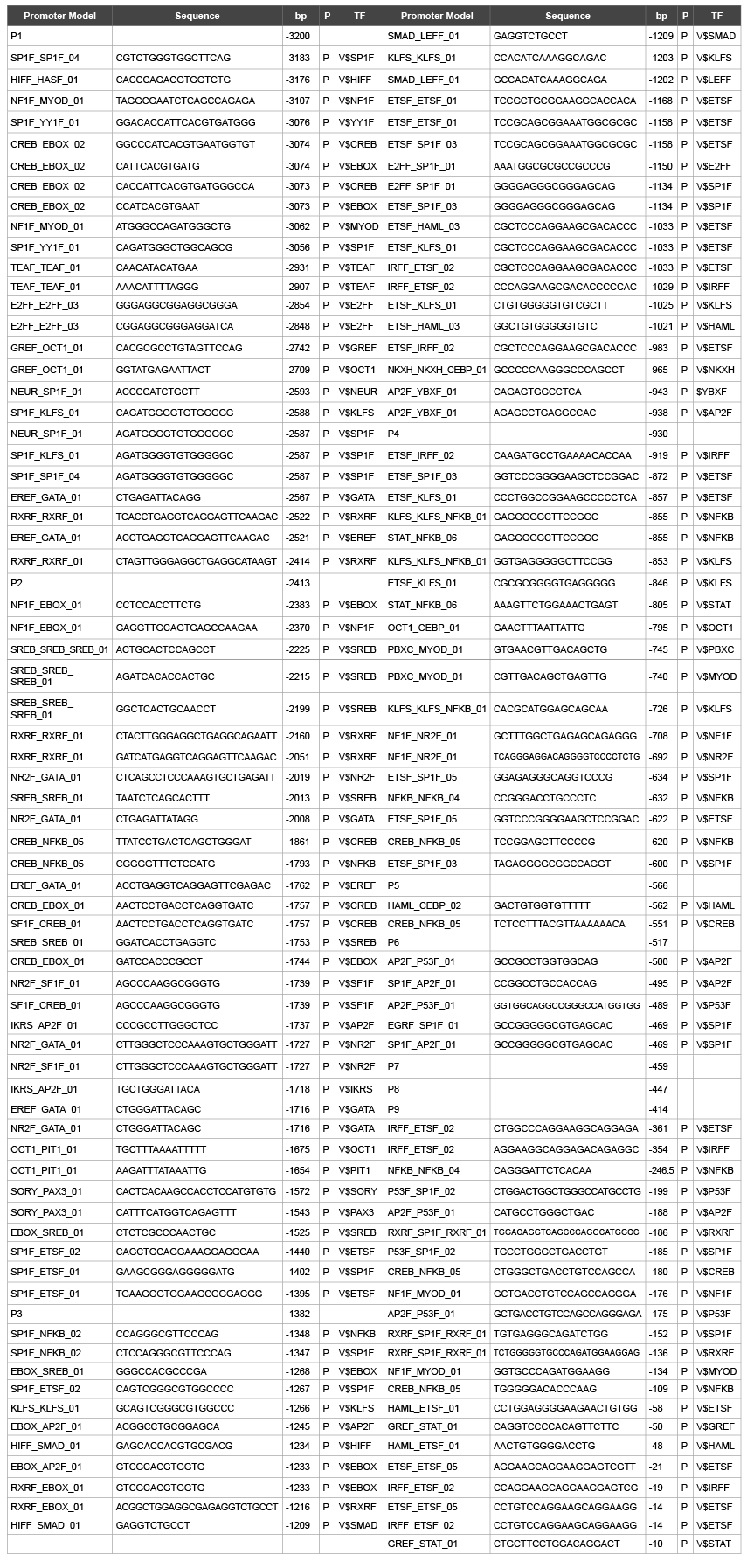
Table 4: Phosphorylation site on promoter model of hMC1R

Table 5: Glycosylation site on promoter model of hMC4R
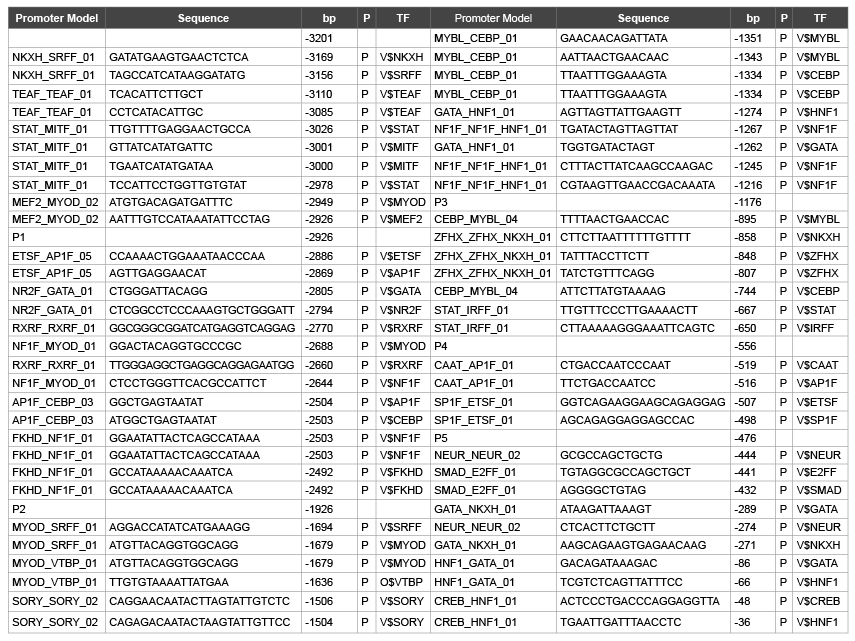
Table 6: Phorsphorylation site on promoter model of hMC4R
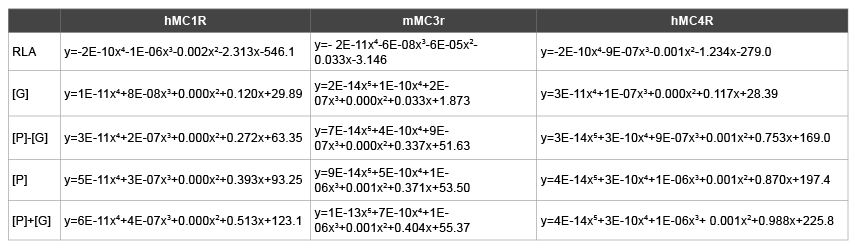
Table 7: Polynormial schemes of trend line RLA, [P], [P] ± [G], [G] for three melanocortin receptors
As a result, for MC1R, trend line of RLA (y=-2E-10x4 -1E- 06x3 -0.002x2 -2.313x-546.1) and trend line of [G] (y=1E-11x4 +8E- 08x3 +0.000x2 +0.120x+29.89) tend to have a relation of negative reciprocal (Figure 3b). At the same time, trend line of RLA (y=-2E-10x4 -1E- 06x3 -0.002x2 -2.313x-546.1) and trend line of [P]-[G] (y=3E-11x4 +2E- 07x3 +0.000x2 +0.272x+63.35) tend to have a relation of reciprocal as well (Figure 3a). For Mc3r, trend line of RLA (y=-2E-11x4 -6E-08x3 - 6E-05x2 -0.033x-3.146) and trend line of [G] (y=2E-14x5 +1E-10x4 +2E- 07x3 +0.000x2 +0.033x+1.873) tend to have a relation of negative reciprocal (Figure 1b). At the same time, trend line of RLA (y=-2E-11x4 -6E- 08x3 -6E-05x2 -0.033x-3.146) and trend line of [P]-[G] (y=7E-14x5 +4E- 10x4 +9E-07x3 +0.000x2 +0.337x+51.63) tend to have a relation of negative reciprocal as well (Figure 1a). For MC4R, trend line of RLA (y=-2E-10x4 - 9E-07x3 -0.001x2 -1.234x-279.0) and trend line of [G] (y=3E-11x4 +1E- 07x3 +0.000x2 +0.117x+28.39) tend to have a relation of negative reciprocal (Figure 4b). At the same time, trend line of RLA (y=-2E-10x4 -9E-07x3 - 0.001x2 -1.234x-279.0) and trend line of [P]-[G] (y=3E-14x5 +3E-10x4 +9E- 07x3 +0.001x2 +0.753x+169.0) tend to have a relation of negative reciprocal as well (Figure 4a).
For all three receptors (MC1R, Mc3r and MC4R), not only the trend lines of RLA with [G] but also those of RLA with ([P]-[G]), tend to have a negative reciprocal relationship. Further investigations are being made to verify the reciprocal relationship.
As indicated from the figures that the trend line of RLA of all three receptors tend to have a relation of negative reciprocal to that the trend line of their [G] respectively (Figures 1b,3b and 4b), and for all of the three receptors, trend line of [G] is always the closest one (among the negative reciprocal relations) to its negative reciprocal RLA (Table 7). If it is true, Change of signal sensing phosphorylation [P] won’t influence the negative reciprocal regulation of transcription activity by [G], therefore, [G] is independent from [P]. It helps us recall the theory: O-GlcNAc and O-phosphate exhibit a complex interplay on signaling, transcriptional, and cytoskeletal regulatory proteins within the cell, sometimes, O-GlcNAcylation and O-phosphorylation appear to be independently regulated [12]. Does this figure give specific digit evidence to “independently regulated” in the above theory?
In contrast, From the fact shown by the figures that trend line of RLA of all three receptors tend to have a relation of negative reciprocal to that the trend line of the corresponding ([P]-[G]) (Figures 1a,3a and 4a), and for all the three receptors, trend line of ([P]-[G]) is the second closest one (among the negative reciprocal relations) which approaching to its corresponding negative reciprocal RLA (Table 7). If it is true, the structure and nutritional element [G] will reduce the regulation of signal sensing phosphorylation [P] to the transcription activity; [P] and [G] coordinately play a negative reciprocal regulation to the transcription activity? Since O-GlcNAc and O-phosphate exhibit a complex interplay on signaling, transcriptional, and cytoskeletal regulatory proteins within the cell, one of the major functions of O-GlcNAc is to prevent O-phosphorylation and, by doing so, to modulate signaling and transcription in response to cellular nutrients or stress [14], does this negative reciprocal between trend line of ([P]-[G]) and that of RLA give a specific digit evidence to “O-GlcNAc prevent O-phosphorylation” in the above theory? Notice here is glycosylation, not O-GlcNAc.
Moreover, for all three receptors, as shown in Table 7, sequence of the trend line is: RLA~[G]<([P]-[G])<[P]<([P]+[G]), that the ([P]+[G]) is the farthest away from the RLA. From opposite aspect, which gives another specific digit proof to the above theory, major function of O-GlcNAc is to prevent O-phosphorylation. Since if [G] play a positive role in the [P], trend line of the ([P]+[G]) would be closer to that of RLA.
Now the question of investigation is does the nutritional and obesity related glycosylation and the signal sensing phosphorylation regulated the transcriptional activity in a negative reciprocal way for the MC1R, Mc3r and MC4R neural receptors? Besides, no matter such negative reciprocal exist or not in experimental world, simply from the negative reciprocal relationship indicated by the RLA ~([G] or [P]-[G]) figure, we can predict the transcriptional activity of the MC1R, Mc3r and MC4R in the same cell type as shown in their corresponding RLA, by counting the [P] and [G].
There is no conflict of interest that could be perceived as prejudicing the impartiality of the research reported.
This research did not receive any specific grant from any funding agency in the public, commercial or not-for-profit sector.
The authors are grateful to all who has helped us.
Download Provisional PDF Here
Article Type: Research Article
Citation: Wen X, Zeng WW (2016) Sweet Regulation of Melanocortin Receptor (MC1R, Mc3r, and MC4R) Transcription Activity. Int J Endocrinol Metab Disord 2(1): doi http://dx.doi.org/10.16966/2380-548X.122
Copyright: © 2016 Wen X, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Publication history:
All Sci Forschen Journals are Open Access