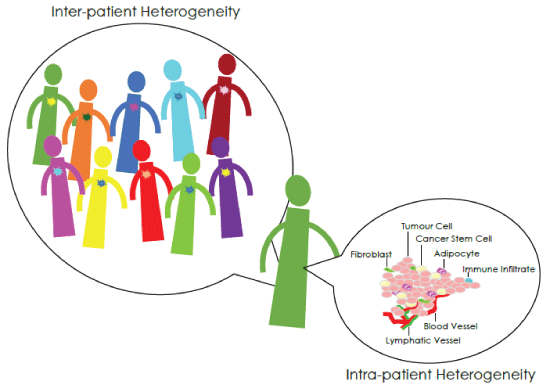
Figure 1: Inter-tumour and Intra-tumour Heterogeneity


Concettina La Motta*
Department of Pharmacy, University of Pisa, Pisa, Italy*Corresponding author: Concettina La Motta, Department of Pharmacy, University of Pisa, Pisa, Italy, Tel: +30 050 2219593; E-mail: concettina.lamotta@unipi.it
Since the pioneering studies of Heppner and co-workers on mouse mammary cancer [1], the presence of heterogeneous phenotypes and genotypes within the cancer bulk has been clearly demonstrated in several types of malignancies [2].
Different co-existing sub-populations, arising from both genetic and non-genetic variability and providing both heritable and non-heritable clones, give rise to an organized community in which not only cooperative but also conflicting interactions may set up. The resulting clonal crosstalk guarantees tumour growth, helping its spread through metastasis and the emergence of resistance to the commonly exploited anti-cancer treatments.
Although aware of tumour heterogeneity [2-6], we are still far from understanding how different phenotypes and genotypes interact, thus allowing the whole community to survive therapeutic selection and/ or suppression. This is why we failed so far to fine-tune a long-lasting, effective treatment for this kind of pathology. Accordingly, untangle tumour complexity is a basic and urgent challenge scientists should not work around anymore.
Different kinds of cancer heterogeneity exist, regarding both intertumour and intra-tumour differences (Figure 1) [7]. Malignancies belonging to the same histopathological subtype but affecting different patients show genetic diversity, and are characterized by divergent functional behaviours. At the same time, malignancy affecting the same patient shows a marked variation too: the tumour bulk harbours both phenotypically and genotypically different cell populations (spatial heterogeneity), and heterogeneity may also arise when serial samples from the same patient are examined within the entire disease course (temporal heterogeneity).

Figure 1: Inter-tumour and Intra-tumour Heterogeneity
A number of players take part to this heterogeneous tumour architecture. The main portion of the tumour bulk is made up of rapidly cycling and terminally differentiated cancer cells, but a small percentage of cells co-exist, bearing stem-like properties, whose origin is still a matter of debate [8-10]. They have been described as either arising from normal stem cells, as a result of epigenetic mutations, or deriving from more differentiated cells, which have acquired self-renewing ability through genetic modifications. As an alternative, Cancer Stem Cells (CSCs) may be generated by an Epithelial-to-Mesenchymal Transition (EMT), which allows the existing neoplastic cells to acquire mesenchymal traits and express stem-cell markers.
According to the hierarchical model of tumorigenesis, the so called ‘cancer stem cell theory’, CSCs play a key role in tumour perpetuating and progression, thanks to their indefinite self-renewal ability and aberrant differentiation capacity [11].
Moreover, differently from the bulk cells, CSCs resist to chemotherapy and radiotherapy. Indeed, cytotoxic treatments commonly exploited to target rapidly dividing cells are rather ineffective against this specific cancer sup-population, as it is characterized by a slow mitotic activity possibly associated with a quiescent G0/G1 state. Therefore, rather than being eradicated, CSCs turn out to be selected over the cancer bulk, being free to drive tumour relapse and mediate metastasis [12-16].
It is now clear that both CSCs and the non-stem differentiated tumor cells exist in a dynamic equilibrium, as the former can differentiate through asymmetric division and the latter can acquire CSC-like state. Any shift in this balance leads to a more CSC-rich tumor, which increases its aggressiveness worsening patients’ prognosis.
The reversible and highly plastic stem-like state is modulated by micro environmental signals arising in the tumor niche [17-20]. Indeed, microenvironment surrounding the tumour bulk and comprising non-neoplastic cells like fibroblasts, adipocytes, infiltrating immune/ inflammatory cells, as well as vessels and associated matrix, does not play a mere role of scaffold but rather participates actively in tumour growth modulation through both stimulatory and inhibitory signalling, thus further ravelling tumour complexity.
Bearing in mind the overall picture, a fruitful treatment of tumour should not ignore a careful selection of patients, made on the analysis of both tumour cells types and microenvironment hallmarks. In particular, the commonly available sequencing technologies should be used at diagnosis, to mark tumour heterogeneity, but also thereafter, to monitor cell differentiation during treatment response and point out the possible emergence of drug resistance during disease progression. Accounting for heterogeneity allows to choose the best personalized therapeutic approach which, in turn, should necessary combine agents targeting cancer’s growth and survival, focusing in particular on CSCs, with derivatives targeting tumour microenvironment. Complementarity in the mechanism of actions of different compounds may merge, in principle, the best functional response with the lowest risk of drug resistance, thus helping to achieve a plain recovery.
Download Provisional PDF Here
Article Type: Editorial
Citation: La Motta C (2017) Unveiling Heterogeneity: The Pressing Challenge to Cancer Diagnosis and Therapy. J Drug Res Dev 4(1): doi http://dx.doi.org/10.16966/2470-1009.e102
Copyright: © 2017 La Motta C. This is an openaccess article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Publication history:
All Sci Forschen Journals are Open Access