Introduction
The advancement of biotherapeutic development demands the application of analytical methods that allow the selection of candidates that provide the best therapeutic value. The affinity of a therapeutic candidate may be closely correlated with its therapeutic value as it has been shown that an increase in affinity of antibody-antigen interaction may result in an increase in potency [1,2]. Therefore a significant amount of effort is devoted to obtain high affinity antibodies [1,3,4], even in cases where an inverse relationship between affinity and activity is observed, as shown in the case of certain agonistic EPOR antibodies [5]. Affinity-based selections of the candidates can be challenging due to the trend towards developing extremely high affinity antibodies, in the picomolar and sub-picomolar range, for many therapeutic areas; therefore, the affinity measurement requires methods suited to high-affinity interactions. It is also important to analyze affinities in physiologically relevant conditions (or complex matrices) as it accounts for the presence of serum components that could potentially alter the interaction of the antibody with its antigen. Analysis under these conditions will also allow the study of the interaction with the antigen in its natural environment and in the presence of factors that could be important for retaining functionality and biophysical properties.
The most widely used techniques for affinity analysis include Biacore and ProteOn (surface plasmon resonance), Octet (biolayer interferometry), KinExA (Kinetic exclusion Assay), ITC (Isothermal titration calorimetry) and Scatchard Aanalysis (radioimunoassay), among other methods [6-12]. These are powerful tools to determine a wide range of affinities (µM to sub-pM), although each has limitations. These include limits in sensitivity or throughput, the need for immobilization or labeling of one of the interactants (such as the use of radioactive material), incompatibility with complex samples matrices or a combination thereof. Several research groups have worked on developing methods to overcome one or more of these limitations. In 2012 Salimi-Moosavi et al. [13] published the development of a label-free, immobilization-free method that retained the sensitivity of the KinExA method and overcame the sensitivity limitations of some of the other methods such as SPR or ITC. This technique is a solution equilibrium affinity (SEA) method using the microfluidic platform of the Gyrolab instrument [14], which improved the analysis throughput by 10-fold in comparison with KinExA [13]. Other efforts were initiated for HTP binding analysis of Fabs using the MSD instrument [15,16]. More recently, Estep et al. [17] refined this concept and published their work on a method using the Meso Scale Discovery (MSD) instrument as a detection platform for solution equilibrium affinity measurements. They demonstrated the applicability of this labelfree method for HTP affinity analysis of antibody candidates, providing a higher throughput than the method reported by Salimi-Moosavi et al. [13]. The method published by Estep et al. [17] is a more sensitive alternative to the method developed by Haenel et al. [18] using the BioVeris instrument (no longer commercially available) or other HTP methods previously described [19]. Measuring affinity in complex matrices such as high concentrations of serum remained one of the most severe limitations of affinity determination until recently, when Bee et al. [20] applied KEA for the analysis of samples in serum [20]. In addition to the studies performed by KEA, others have shown advances using backscattering interferometry [21,22] for working with neat serum (personal communication). In this work we describe a method that combines the high throughput capability of MSD to determine SEA as demonstrated by Estep et al. with tolerance to neat serum and demonstrate its application to the analysis of carlumab binding to CCL2. CCL2 (CC-chemokine ligand 2), is a chemokine which induces calcium mobilization and chemotaxis upon binding to its receptor (CCR2, a G-protein-coupled receptor) [23]. CCL2 has been implicated in inflammatory diseases [24] and cancer [25-28]. Carlumab is a human antibody that entered clinical trials for oncology [29]. It was noted in the trial that carlumab briefly suppressed free CCL2, but this effect was not sustained with dosing. Due to lack of single agent efficacy, development of the drug was discontinued. One of the approaches taken to investigate these observations was to compare the affinity of the antibody in PBS versus neat sera to determine if the interaction of carlumab with CCL2 was affected by the presence of serum. In this exercise it was important to keep serum as unaltered as possible to perform the analysis in the most relevant conditions.
We report here the affinity determination for carlumab in human sera and the method we developed to accomplish this goal. We compared the results obtained with the MSD-SEA (solution equilibrium affinity analysis using Meso Scale discovery technology) method with data obtained by surface plasmon resonance analysis (Biacore and ProteOn), kinetic exclusion assay (KinExA) and isothermal titration calorimetry (ITC). The MSD-SEA experiments were performed in PBS or in neat human sera. The affinity results obtained in PBS by Biacore, KEA and ITC were in agreement with the results obtained by MSD-SEA. Importantly, the results obtained in human sera were in agreement with the results obtained in PBS, demonstrating that carlumab retained high affinity for CCL2 in either matrix, and that lack of efficacy was not due to reduced affinity in serum. This work demonstrates the use of MSD-technology as a detection platform for solution equilibrium affinity determinations in complex matrices, making it possible to analyze interactions in a physiologically relevant fluid, in an HTP format and without the need for labeling of the reactants
Materials and Methods
Materials
Monoclonal antibody against human CCL2 (carlumab) was generated as described by Carton and collaborators [30] and prepared in house (Centocor R&D ) using different cell lines. The antigen binding fragment (Fab) for carlumab was generated as previously described [30-32]. Recombinant CCL2, with BSA as carrier (or carrier free for KEA beads preparation and biotinylation) was obtained from R&D Systems Inc. (cat# 279-MC/CF). Affinity purified indodicarbocyanine (cy5)-labeled goat anti-human (H+L), and goat anti-human IgG Fcγ fragment specific antibody (catalog No. 109-005-098) were obtained from Jackson Immuno Research laboratories Inc. Anti-idiotypic (anti-id) were prepared and labeled in house (Centocor R&D). CNTO402 (C1767A, human IgG1 anti-id for carlumab), was biotinylated and CNTO6707 (C1766, human IgG1 anti-id for carlumab) was ruthenium labelled. MSD Read Buffer (Cat # R92TC-1) and MSD Streptavidin Standard plate (Cat # L11SA-1) were obtained from Meso Scale Discovery. Azlactone activated beads (Cat# 53111) and SuperBlock® blocking buffer in PBS (cat# 37515) were obtained from Pierce Biotechnology. Sensor chips amine, coupling reagents, and surfactant P20, were obtained from (Biacore AB; currently part of GE Healthcare) or from Bio-Rad. Dulbecco’s phosphate-buffered saline (PBS), pH7.4, from Invitrogen Corporation (cat# 14190). Human serum was obtained from Bioreclamation Inc. IgG1 from human myeloma plasma was obtained from Sigma (cat# I5154).Other reagents were, bovine serum albumin, fraction V (Sigma, A3059-100G), Tween 20 (Sigma, P1379). MSD Assay and blocking buffer: 10 mM phosphate buffered pH 7.4, 150 mM NaCl, 1 mg/mL (1%) BSA, 0.05% Tween 20. MSD wash buffer: 10 mM phosphate buffered pH 7.4, 150 mM NaCl, 0.05% Tween 20. MSD Read Buffer (Meso Scale Discovery, R92TC-1), MSD Streptavidin Standard plate (MSD, L11SA-1) were obtained from Meso Scale Discovery. Other equipment and consumables was a Tecan plate washer or BIO-TEK automated plate washer with stacker, sample plate (Greiner 651201, 0.3 mL v-bottom).
Surface plasmon resonance (SPR) experiments by Biacore in
PBS buffer
Surface plasmon resonance experiments were performed using a Biacore 3000 optical biosensor (Biacore AB; currently part of GE Healthcare). The biosensor surface was prepared by coupling goat anti-human IgG Fcγ fragment specific antibody to the carboxymethylated dextran surface of a CM-5 sensor chip using the manufacturer’s instructions for aminecoupling chemistry. The coupling buffer was 10 mM Na acetate, pH 4.5. About 1800 RU of Ab was captured in each reaction and reference flow cells.
Kinetic experiments were performed at 25°C in PBS (containing 0.005% surfactant P20 and 3 mM EDTA). PBS was filtered and degassed prior to use. Five serial dilutions of CCL2 were prepared ranging from 5 nM to 60 pM. About 95 RU of carlumab were captured on flow cell 2 or flow cell 4 using carlumab at 1 µg/mL. A flow cell modified with anti-Fc antibody but lacking carlumab was used as reference. Capture of carlumab was followed by injection of 240 µL (association phase) of CCL2 at 60 µL/min, followed by 900 or 1800 seconds of buffer flow (dissociation phase). The chip surface was regenerated by a 20 µL injection of 100 mM H3PO4 followed by a 20 µL injection of 50 mM NaOH, at 60 µL/min. For data processing, double reference subtraction of the data was performed to correct for buffer contribution to the signal and instrument noise [33] using the Scrubber software version 1.1 g (BioLogic Software) for referencing. After this initial data processing, kinetic analysis of the data was performed using the BIA evaluation software, version 4.0.1 (Biacore, AB) using a Langmuir 1:1 binding model.
Surface plasmon resonance (SPR) experiments in serum by
Biacore and ProteOn
SPR experiments in serum were performed using a Biacore or a ProteOn XPR36 instrument (Bio-Rad). Two types of biosensor surfaces were prepared. The first type of surface was prepared by direct covalent immobilization of carlumab onto the sensor surface. The second surface was prepared by covalent immobilization of goat anti-human IgG Fcγ fragment specific antibody to the surface of a CM-5 chip (for Biacore) or a GLC sensor chip (for ProteOn) using the manufacturer’s instructions for amine-coupling chemistry. The coupling was performed following the conditions described in the previous section.
The trials to perform kinetic experiments were conducted at 25°C using running buffer (PBS pH 7.4 containing 0.005% surfactant P20, and 3 mM EDTA). CCL2 was prepared in running buffer or neat serum. The running buffer was filtered and degassed prior to use. For the surface modified with anti-Fc antibody, about 95 RU of carlumab were captured in one of the flow cells or channels. Carlumab capture was followed by injection of human IgG from myeloma plasma (Sigma, cat # I5154). CCL2 was injected over the sensor chip containing captured or covalently immobilized carlumab. The association was monitored by injection of 240 µL of 5 nM CCL2 at 60 µL/min and the dissociation was monitored for 1800 seconds (of buffer flow). The chip surface was regenerated by a 20 µL injection of 0.85% H3PO4 , at 100 µL/min. Data processing was performed using the manufacture’s software for ProteOn or Scrubber software version 1.1g (BioLogic Software).
Isothermal titration calorimetry
Isothermal titration calorimetry (ITC) experiments were performed using a VP-ITC micro calorimeter (Microcal Inc.) at 20.8°C in 20 mM MES buffer, pH 6.0, containing 100 mM NaCl. Titrations were performed by injecting 18 µL aliquots of 20 µM CCL2 into 2 µM solution of carlumab Fab. The area under each injection was integrated to obtain the heat of reaction. Heats of reactions were corrected for the effect of dilution. The instrument was calibrated using standard procedures as recommended by the manufacturer, and by the titration of 1 mM HCl into 10 mM Trisbase (Sigma, Co). Data were analyzed using the Origin software, V7.083 (Origin Lab Corporation) and fitted to a one site binding model.
Kinetic exclusion assay (KEA)
Kinetic exclusion assays were performed using a KinExA 3000 instrument (Sapidyne Instruments). Equilibrium experiments were conducted at RT and using a 1:1 dilution of PBS and superblock blocking buffer (in PBS) as buffer. The working concentrations of CCL2 were 766.7, 187.0, 47.9, 7.48, 0.75, and 0.037 pM, while the working concentration of carlumab was 15 pM. Antigen/antibody samples were allowed to reach equilibrium by incubation of the mixtures for 4 h at 37°C, followed by 13 h incubation at RT. To detect free carlumab the equilibrated samples were loaded onto the KinExA 3000 flow cell which had been prepared with a column of CCL2-modified azlactone activated beads. Cy5 labeled goat anti-human (H+L) antibody was used for detection of free carlumab which bound to the beads in the equilibrated samples. Experiments were performed in duplicate. The data was fitted to a 1:1 binding model using the KinExA Pro software, version 1.0.3 (Sapidyne Instruments).
MSD-SEA experiments in PBS vs serum
Solution equilibrium affinity determinations using the MSD instrument for detection (MSD-SEA method) were performed by diluting the samples in buffer or in neat serum by preparing the complexes in a similar manner as they were prepared for KEA experiments using the KinExA instrument. However, the detection of free mAb was performed using an MSD reader instrument (MSD Sector Imager 6000TM Reader).
A serial dilution of CCL2 in neat serum or buffer (10 nM to 0.01 pM in 4-fold dilution series) was prepared in 96-well polypropylene sample plates. A negative control containing buffer or serum without antibody or antigen and a positive control containing antibody without antigen were also included. When the samples were prepared in buffer, the buffer (MSD Assay and blocking buffer) consisted of 10 mM phosphate buffer pH 7.4, 150 mM NaCl, 1 mg/mL 1% BSA, 0.05% Tween 20. Carlumab at 20, 40 or 60 pM was also prepared in neat serum or buffer. Equal volumes (50 µL each) of each constant concentration of carlumab were mixed to each CCL2 concentration; therefore, the final concentrations were half of those indicated above. The samples were mixed by shaking the plates briefly and reached equilibrium by incubation with constant rotation for 24 or 48 hours at 4°C. Duplicates of the mixtures were incubated with or without constant rotation. After incubation, unbound (free) carlumab in the equilibrated mixture was detected by Electro Chemiluminescene Immunoassays (ECL).
For detection of free carlumab detection plates were prepared the day prior to analysis. The plates were prepared by blocking MSD Streptavidin Standard plates with assay buffer for 5 minutes, followed by addition of 200 ng/mL of biotinylated CNTO402 (C1767A; a neutralizing antiidiotypic antibody (anti-id) against carlumab in assay buffer followed by incubation for ~18 hours at 4°C. At the time of detection the plates were blocked by adding 150 µL/well of assay buffer without removing coating capture reagent and incubated for 1 hour at ambient temperature. This was followed by 5X washing with wash buffer. After this 50 µL/well aliquot of the carlumab/CCL2 mixture were transferred to the detection plate and incubated for 30 minutes to capture free carlumab onto biotinylated CNTO402 (C1767A)), followed by 3 times washing with Wash Buffer. Next 50 μL per well of 0.26 µg/mL of ruthenium labeled CNTO6707 (C1766A) were added and incubate for 1 hour. The plates were washed and 150 µL per well of MSD Read Buffer (1:4 of stock into deionized H2O) were added. The plates were read immediately on the MSD Sector Imager 6000TM Reader for detection of luminescence levels. The data was analyzed using a GraphPad Prism software version 5 (GraphPad Software Inc.). The raw data was normalized using Prism normalizing function to define the curve from 0% to 100% free antibody using buffer or neat serum control signal (no carlumab, no antigen ) as the 100% antibody bound. The normalized data was fitted to a non-linear regression analysis curve to determine the affinity of the antibody for the antigen. A user defined equation was entered into Prism which was generated from the law of mass action for dynamic equilibrium [18,34-36] originally described by Guldberg & Waage [36], and revisited for several systems by others [18, 34-36].
Results
Kinetic analysis in buffer using Biacore
SPR experiments using Biacore were performed to determine the affinity of carlumab to CCL2 in PBS buffer (PBS, 0.005% P20 and 3 mM EDTA, pH 7.4). Figure 1 shows a typical Biacore binding profile for carlumab including dissociation data collected for 30 minutes providing at least 5% (>20%) response decay which is needed for accurate determination of the dissociation rate constant [37]. Samples of carlumab from five different expression systems were analyzed. Carlumab from the cell line SP2/0 was used as a control to account for any potential differences between flow cell two and flow cell four. Table 1 summarizes the results for the affinities (KD), and the stabilities (kd) of the complexes formed by the five different samples of carlumab and CCL2 as determined by Biacore. KD values range from 23 to 38 pM which shows tighter average affinities for the samples expressed in CHO-C1463A as compared to the other 4 cell lines, specially SP2/0. However, the standard deviation and comparison of the data for SP2/0 in flow cell 2 versus flow cell 4, suggest that there may not be significant differences between the affinities of carlumab produced in these cell lines. These results indicate that carlumab binds to CCL2 in PBS with high affinity and that the affinity is minimally affected by the differences in the expression system.
Carlumab cell line |
Flow cell |
ka (M-1s-1)
|
kd(s-1) |
KD(pM)
|
CHO-C1463A |
2 |
(10.1 ± 0.10) × 106 |
(2.47 ± 0.68) × 10-4 |
25 ± 7 |
CHO-C1493A |
4 |
(9.09 ± 0.99) × 106 |
(2.58 ± 0.15) × 10-4 |
28 ± 4 |
CHO-C1494A |
2 |
(9.14 ± 0.27) × 106 |
(2.08 ± 0.38) × 10-4 |
23 ± 8 |
HEK-293 |
4 |
(9.21 ± 0.75) × 106 |
(2.52 ± 1.11) × 10-4 |
38 ± 12 |
SP2/0 |
2 |
(10.1 ± 0.17) × 106 |
(2.81 ± 0.54) × 10-4 |
28 ± 7 |
SP2/0 |
4 |
(7.76 ± 0.13) × 106 |
(2.97 ± 0.79) × 10-4 |
38 ± 12 |
Table 1. Biacore results: Kinetic and affinity data for the interaction of carlumab with CCL2. The sample carlumab SP2/0 was captured in Flow cell 2 and flow cell 4 to account for any differences due to the sequential flow of the sample through the flow cells. Shown are the average parameters with the corresponding standard deviation for three different e×periments with samples injected in triplicate in most cases. Two sensor chips were used.
Isothermal titration calorimetry (ITC) confirms SPR results
and defines stoichiometry for carlumab-CCL2 interaction in
buffer
Isothermal titration calorimetry (ITC) was carried out to characterize the binding of carlumab to CCL2, to gain insights into the forces driving the interaction and to confirm the stoichiometry estimated by Biacore. ITC is not usually the choice method to determine affinity in the picomolar range; however, it is the optimal technique for determination of stoichiometry. The ITC data revealed that carlumab Fab bound to CCL2 with 1:1 stoichiometry of ligand:Fab (Figure 1b) corroborating the stoichiometry obtained by Biacore. Although the binding interaction exceeded the ITC limits for accurate measurements of affinity, as indicated by a very sharp transition in the binding curve, the results corroborate that the affinity is sub-nanomolar. In addition, direct and accurate measurements of the change in enthalpy (ΔH) can be obtained by ITC [38]. A negative change in enthalpy of -7.9 ± 3.5 kcal/mole of CCL2 was observed and shows that enthalpy contributes to driving the interaction at 25°C. It has been shown that enthalpic contributions originate from changes in protein-protein, protein-solvent and solvent-solvent H-bond network, formation of salt bridges, and changes in van der Waals contacts [39]. Negative ΔH has been attributed mainly to specific interactions such as H-bonds and salt bridges [39]. The data presented here suggest that these kind of molecular forces drive the interaction between carlumab and CCL2. This is in agreement with the results obtained by X-ray crystallography, which revealed that basic residues such as arginine are involved in the complex formation, forming several hydrogen bonds between the antibody and the antigen [31]. The antigen-antibody interphase also includes van-der-Waals interactions [31], that together with salt bridges and hydrogen bonds are the forces driving the binding leading to a negative ΔH.
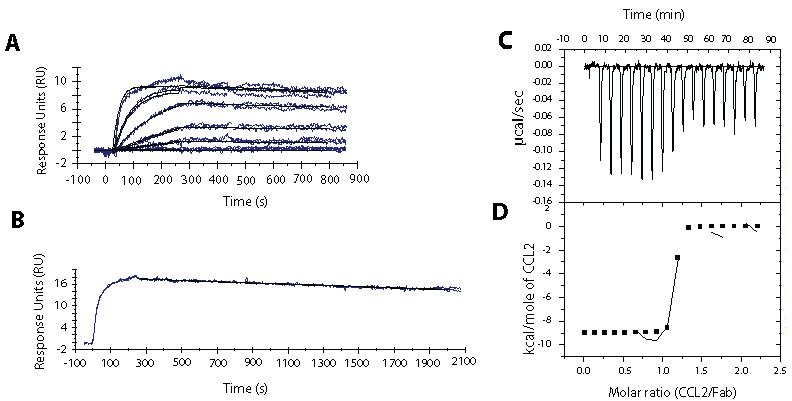
Figure 1: Biacore and ITC experiments. Typical Biacore sensorgrams for the kinetic titration of carlumab with human CCL2 (A), and the dissociation phase data collected for 30 minutes (B). The solid smooth lines correspond to the fit to a simple 1:1 binding model. ITC data showing heats of binding of CCL2 to the Fab of carlumab (C) and the heats of reaction normalized to the number of moles of CCL2 injected and are corrected for the heat of dilution (D). The solid line is the nonlinear least-square fitting the data to a single site model.
Label-free solution equilibrium experiments by KEA confirms
SPR results for the antigen-antibody interaction in buffer
In order to confirm the affinities obtained by SPR, we performed solution equilibrium affinity (SEA) analysis using the kinetic exclusion assay (KEA) method. KinExA has been recognized as an enabling technology that allows the study of high affinity interactions (pM and sub-pM) including those with very fast on-rates (>107M-1s-1) and offrates slower than 10-5s-1 [8]. KEA is a highly sensitive fluorescence-based method and it has been described in detail by Darling and Brault and others [8,9]. In order to measure the affinity of carlumab with CCL2 in solution, we prepared mixtures containing a fixed concentration of carlumab and a serial dilution of CCL2. The mixtures were incubated for greater than 12 hours to reach equilibrium. Free carlumab was detected by flowing the incubated mixture through a KinExA cell packed with CCL2-modified beads and detecting bead-bound carlumab (free carlumab in the reaction mixture) using a fluorescently labeled anti-human antibody. Table 2 (KD) show the KEA results for the complexes formed by five different samples of carlumab and CCL2. The KEA data confirms the results obtained by Biacore with affinities between 17 pM and 63 pM. As seen in Biacore, there is an apparent trend for 2-fold tighter affinity for carlumab expressed in CHO-C1463A compared to carlumab expressed in the other cell lines.
Carlumab cell line |
KD (pM)
|
95% confidence interval |
CHO-C1463A |
17 |
5-36 |
CHO-C1493A |
63 |
45-86 |
CHO-C1494A |
12 |
8-15 |
HEK-293 |
40 |
32-48 |
SP2/0 |
38 |
35-41 |
Table 2. Equilibrium dissociation constants for the interaction of carlumab with CCL2 as obtained by KEA
Kinetic analysis in serum using Biacore and ProteOn
SPR experiments (using the Biacore or ProteOn technologies) were performed with the intention to determine whether the kinetics and affinity of carlumab binding to CCL2 were altered by the presence of serum components. Performing these experiments in neat serum was expected to complicate SPR analysis due to increased nonspecific protein binding and bulk reference subtraction issues. However, since SPR offered higher throughput and has been demonstrated to work with lower concentrations of serum [40,41] developing an SPR method was considered a worthy effort. To better mimic the conditions used in Biacore for the experiments in PBS described above, our serum experiment using Biacore involved capture of carlumab with anti-Fc antibodies. One difficulty with this approach was the presence of human immunoglobulins in serum; therefore, the antihuman Fc-modified sensor surfaces were blocked by injecting saturating levels of purified serum IgG1 antibodies following capture of carlumab. An additional question to the potential effect of serum components in the interaction was whether high concentrations of carlumab (as in the case of high patient dosing) could alter the interaction. This was counterintuitive given what it is known of the molecular interactions with immobilized molecules on a sensor surface; however, it was a valid question as the potential effect of serum on the affinity of carlumab came into question from observations of more free CCL2 than expected with continuous and increasing dosing of carlumab [29]. To answer this question 3 different level of carlumab capture ranging from 16 to 620 RU were used. The effect of the different RU levels of carlumab was tested first in PBS by injecting 5 nM CCL2 and observing the dissociation for 30 minutes. The data revealed that in PBS the increase in surface concentration of carlumab did not result in an increase in dissociation rate (data not shown). At 159 RU, 321 RU and 618RU of carlumab immobilized onto the surface the dissociation rate was 2.79 x 10-4s-1, 1.49 x 10-4s-1 and 7.70 × 10-5s-1 and 7.70 (7) × 10-5s-1 respectively, indicating a trend towards stabilization of the interaction and not a weakening of the interaction. After this the experiment was repeated by injecting 5 nM CCL2 in neat serum. This experiment in Biacore was unsuccessfully due to instrument clogging and high nonspecific binding. The experiments were continued using the ProteOn instrument, which offered a higher throughput (where different conditions could be tested simultaneously), different fluidics and different sensor matrices. In the ProteOn, we tested carlumab capture via Fc simultaneously with covalently immobilized carlumab by using different spots on the sensor surface.
To prepare the ProteOn sensor surface anti-human Fc antibody was immobilized (~ 6000 RU) on the horizontal orientation (A4-A6) and carlumab was immobilized (35 to 6000 RU) on the vertical orientation (L4- L6) which resulted in a mixture of anti-Fc and carlumab on (A4-A6 x L4-L6). While these mixed spots could not be used we also obtained 6 blank spots in (A1-A3 x L1-L3) that could be used as references (Figure 2a). Carlumab was captured (100-400 RU) on two of the anti-Fc surfaces and one was keep for referencing. To improve the conditions used in Biacore by controlling for nonspecific binding of serum components and to avoid pre-saturation of the carlumab surface with any endogenous CCL2, carlumab Fab was added at 100 nM (a concentration >>KD in PBS) to a portion of the nonCCL2 spiked serum and incubated ~30 min. This serum preparation served several functions, to saturate the surface with serum components (IgG and others) prior to injection of CCL2-spiked serum, to avoid presaturation of the carlumab surface with any endogenous CCL2, as control for double reference subtraction, and to capture a non-relevant antibody on the reference surfaces for better referencing.
The amount of covalently immobilized carlumab varied from ~35 to ~6000 RU (from the lowest captures normally used on SPR to maximum binding capacity of the chip in this case). 6000 RU was ~75 times higher than the capture normally used on Biacore for experiments in PBS.
After carlumab capture and treatment of the sensor chip with carlumab-treated serum, 5 nM CCL2 was injected (observed saturating concentration in Biacore experiment) in neat serum. This injection was followed by monitoring of the dissociation phase in PBS running buffer for 30 minutes. The data showed that the off-rates of CCL2 bound to carlumab are comparable in PBS versus human serum (2.02 × 10-4s-1; Figure 2B). Therefore, we concluded that a change in dissociation rate was not a contributing factor to a potential decrease in binding of carlumab to CCL2 in serum compared to PBS. The possibility remained that there is a difference in the affinity arising from differences in on-rate. However, the association phase could not be used to determine association in these experiments due to ineffective referencing as shown in the portion labeled “assn” in figure 2B. Although it could be concluded that serum had no effect in the dissociation rate the for carlumab-CCL2 interaction, the question remained unanswered for the association rate.
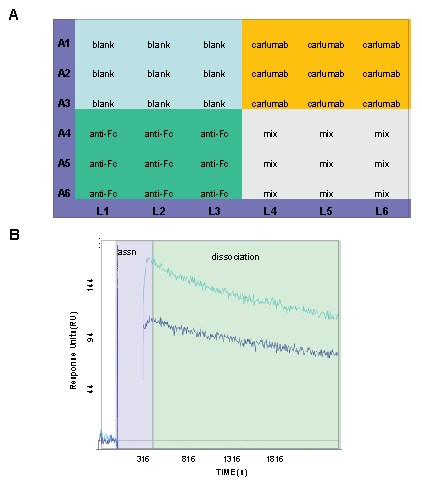
Figure 2: ProteOn in neat serum. (A) ProteOn sensor surface diagram indicating anti-human Fc antibody on the horizontal orientation (A4-A6), carlumab immobilized on the vertical orientation (L4-L6), a mixture of anti-Fc and carlumab on (A4-A6 x L4-L6) which were not usable, and blank spots in (A1-A3 x L1-L3) to be used as reference for covalently immobilized carlumab. (B) Sensorgrams for the injection of 5 nM CCL2 in neat serum over 2 different surfaces (“assn” phase) followed by monitoring of the dissociation phase. These sensorgrams were obtained after controlling for nonspecific binding of serum components by injecting neat serum where carlumab Fab was added at 100 nM (to avoid pre-saturation of the carlumab surface with any endogenous CCL2 carlumab) to non CCL2-spiked serum.
MSD-SEA shows that the affinity of carlumab to CCL2 is
retained in neat serum
In agreement with recent publications [17] our work showed that the MSD platform can be used for the analysis of high affinity interactions in buffer. In order to determine if serum components affect the affinity of carlumab to CCL2, solution equilibrium affinity analyses were performed using the MSD reader as the detection platform using the (MSD-SEA) method. To perform the analysis, the complexes were prepared in a similar manner to the procedure used for KEA but the detection was performed using an electrochemiluminescence assay (ECL) with anti-idiotypic antibodies as capture and detection reagents. The volumes used were 50 µL per sample for a total of 100 µL of reaction mixture. For the detection of free carlumab after equilibration of the reaction mixture, a 50 µL aliquot was transferred to streptavidin (SA) MSD plates which had been coated with biotinylated anti-id to capture free carlumab. Better results were obtained when these detection plates were blocked with assay buffer and incubated with the capture anti-id for ~18 hours at 4°C and blocked with assay buffer (which contained BSA) prior to use for capturing free carlumab. Captured carlumab was detected using a different anti-id as depicted in figure 3A. Free carlumab was captured and after this step, ruthenium-labeled anti-id was added and incubated for 1 hour, followed by washing and detection using the MSD Sector Imager 6000TM Reader.
To fit the data for affinity, we introduced a user defined equation into GraphPad Prism software. This user defined equation was generated from the law of mass action for dynamic equilibrium originally described in [34], and revisited for several systems by others [18,35,36]. Prior to fitting for affinity, the raw data was normalized using Prism normalizing function to define the curve from 0% to 100% free antibody using buffer or neat sera control signal (no carlumab, no antigen added) as the 100% antibody bound. The normalized data was fitted using a non-linear regression analysis curve fit using the user defined equation described by Haenel and collaborators and Piehler and collaborators [18,36]. The signal for carlumab in neat sera with no added recombinant CCL2 was at the level of the plateau reached for 100% free carlumab, thereby indicating no detectable effect of endogenous CCL2 in the sera on the binding of carlumab. Figures 3B and 3C show the binding profiles for carlumab binding to CCL2 in PBS buffer and in neat serum while table 3 shows the KD as determined by MSD-SEA. In addition to the affinities, table 3 shows the parameters that were used to evaluate the goodness of the fit. The table shows R square values >0.98 and standard errors of <10% indicating that the data is very well described by the user defined equation. Finally, the standard deviation of three measurements is 12% and 22% for PBS and neat serum, respectively, indicating good reproducibility of the data in both PBS and serum. The MSD-SEA data (Table 3) is in very good agreement with the KinExA and Biacore data (Tables 1 and 2) obtained in PBS buffer. Importantly, the MSD-SEA data show that the affinity of carlumab to CCL2 in neat human sera is the same as the affinity in PBS, ~40 pM.
Sample Matri× |
KD (pM)
|
R square |
Standard error |
95% confidence interval |
PBS |
48.1 |
>0.98 |
9.5 |
29-67 |
PBS |
42.5 |
>0.98 |
3.9 |
35-51 |
PBS |
37.3 |
>0.98 |
6.4 |
25-50 |
Average |
42.6 ± 5.4 |
|
|
|
Serum |
51.9 |
>0.98 |
9.7 |
33-72 |
Serum |
45.5 |
>0.98 |
6.8 |
32- 59 |
Serum |
32.6 |
>0.98 |
7.5 |
18-48 |
Average |
43.3 ± 9.8 |
|
|
|
Table 3. Equilibrium dissociation constants for the interaction of carlumab with CCL2 as obtained by MSD-SEA.
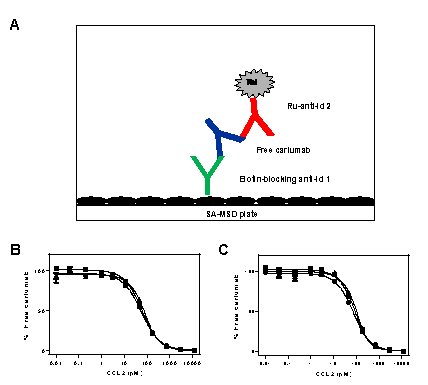
Figure 3: MSD-SEA. Schematic representation of free antibody detection in solution equilibrium affinity experiments by MSD-SEA (A). This diagram depicts the detection of free carlumab in equilibrated carlumab/CCL2 mixtures. Free carlumab in the equilibrated mixture was captured using a neutralizing biotin labeled anti-id which has been immobilized onto an SA plate. Capture of carlumab was followed by detection of the captured antibody by another anti-id. MSD-SEA equilibrium binding profiles for the interaction of carlumab with human CCL2 (B) in PBS and (C) in neat human sera. The solid smooth lines correspond to the fit to a simple 1:1 binding model. The data was obtained from a mixture of a 4-fold serial dilution of human CCL2 (final concentrations of 5 nM to 0.005 pM) and constant concentrations of carlumab (final concentrations of 10, 20 or 30 pM).
Discussion
We have developed a high throughput method using MSD technology (MSD-SEA) that is suitable for solution equilibrium affinity determination in complex matrices, such as neat serum. This method enabled us to demonstrate that the affinity of carlumab for CCL2 was high (~40 pM) and was unaffected by the presence of serum components. The MSD-SEA data showed very good agreement with SPR and KEA and demonstrated that the serum components did not affect detection of the interaction.
The MSD-SEA method described here is designed to determine the equilibrium affinity constant of a mAb, Fab, receptor, protein or peptide (interactant A) for a soluble ligand, receptor or antigen (interactant B) in a complex matrix and in a high throughput format. In summary, the application of this method consisted of four major steps. (I) Reactants were prepared in neat serum and a constant concentration of interactant A was mixed with a serial dilution of interactant B. (II) The mixture was incubated to reach equilibrium. (III) Following this incubation the unbound (free) mAb/Fab in the mixture was quantified using electrochemiluminescence (ECL) using the Meso Scale discovery (MSD) instrument for detection (IV) and Data was then fitted to an equation derived from the law of mass action as described elsewhere to obtain affinity constants. The key for the application of this method was the use of capture and detection reagents in step III that are specific for the free form of reactant A and do not bind to any of the components of the sample matrix (e.g. serum) or interactant B. The generation of anti-id reagents is an essential component of antibody drug development; therefore, this method could be widely applicable in the biotech industry.
In the selection of the capture and detection reagents (e.g. carlumab anti-id) it is important to consider the factors that will determine the suitability of these reagents. One of these factors is specificity. Both reagents must bind the drug specifically and have no interaction with other components of the matrix. A second factor is the inability of the capture reagent to bind the antigen-antibody complex; therefore the capture reagent must be neutralizing, or sterically-hindered from binding the complex. A third factor is the role that the affinity (and kinetics) of the anti-id selected could play in the suitability of the anti-id for a MSD-SEA assay. The apparent affinities measured in a bivalent format for the antiid used here, CNTO402 and CNTO6707, were in the pM and nM range, respectively (data not shown). Although we did not systematically test the effect of anti-id of different affinities we estimate that it would be optimal to use high affinity capture anti-id for free mAb detection to prevent drug dissociation during the plate washing and capture steps.
Upon the CCL2/carlumab complex formation and establishment of a dynamic equilibrium, the free antibody present within the equilibrated mixture is the free mAb that is quantified in the detection step of this assay. It is important to note that the free antibody measured with the MSD-SEA method may not represent the “free drug” measured during in vivo studies. The complexity of the binding events in an in vivo setting is far greater than the situation of an in vitro MSD-SEA assay. This complexity, in combination with the formation of high amounts of antigen-antibody complex, has been shown to lead to discrepancies between in vitro quantification of different species (free, bound or total) of interactants and in vivo results [42,43]. In addition, in an in vivo study, the serum concentration of drug is far higher than the concentrations used in an experiment for the determination of affinity where we strive to work under KD-controlled conditions for the determination of true intrinsic affinities [7]. Pienta and collaborators [29] report that during carlumab clinical trials, drug steady-state serum concentrations above 10 µg/mL (>67 nM) were achieved. This concentration is >1000-fold the intrinsic KD of carlumab to CCL2, therefore, high above the sweet spot for affinity determination described by Bee and collaborators [7] and could lead to conclusions related to in vivo affinity that are not representative of the true affinity. However, we must also consider the possibility that recombinant CCL2 may not be representative of endogenous CCL2 due to potential effects of glycosylation differences, addition of affinity purification tags or other modifications to the recombinant protein. In the case of CCL2, endogenous levels have been determined to be in the 14-79 picomolar levels in humans (internal data, not shown). We did not determine the actual concentration of endogenous CCL2 in the sera used in these studies. However, the signal obtained for carlumab in sera with no spiked CCL2 was at the level of the plateau where carlumab is 100% free in the equilibrated mixture. Therefore, we estimate that the levels of endogenous CCL2 in the sera used in these studies did not affect the results observed in this report.
Although this method has been developed using the MSD reader as a detection platform, it could be applied to other detection systems that allow for HTP analysis and provide appropriate sensitivity and selective detection. The MSD-SEA method can be applied to other biological systems as long as specific detection reagents are available, such as specific capture and detection reagents that will enable capture of only free antibody (or receptor) out of the equilibrated mixture and specific detection of the captured antibody (or receptor) in the midst of the complex mixture. This method has the added advantages that it uses small volumes (100 or less µL/dilution), small amounts of samples and reagents, no radioactive or fluorescent labeling of the interactants is needed and it can be used for crude samples as long as the concentration of one of the reactants is known. If the concentration of the constant partner is not known, solution equilibrium titrations could be performed using different dilution factors of the sample until a hill slope around (-1), which is the ideal slope for a binding isotherm, is obtained (or +1 if the data is plotted in terms of % bound). The time required for affinity determination is significantly reduced with MSD-SEA as compared with other methods, and can be applied to high-affinity interactions in the pM and potentially sub-pM range. A potential limitation could be the unavailability of adequate capture and detection reagents. However, for therapeutic antibodies this should not be a limiting factor since the generation of anti-id reagents is an essential part of development. MSD-SEA could become an integral part of the biotherapeutics candidate characterization and selection process because it could be used to assess the stability of the candidate in a serum matrix and used to predict the functional stability of the candidate when exposed to physiological conditions.
Acknowledgments
This study was sponsored by Janssen Research & Development. We thank Dr. Gabriela Canziani and Dr. Bonnie Wu for providing information on the affinity and neutralizing activity of the anti-ids.




